
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 25,679,769 – 25,679,859 |
| Length | 90 |
| Max. P | 0.591947 |

| Location | 25,679,769 – 25,679,859 |
|---|---|
| Length | 90 |
| Sequences | 10 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 65.79 |
| Shannon entropy | 0.73662 |
| G+C content | 0.46639 |
| Mean single sequence MFE | -20.36 |
| Consensus MFE | -7.75 |
| Energy contribution | -8.25 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.38 |
| Mean z-score | -0.95 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.591947 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
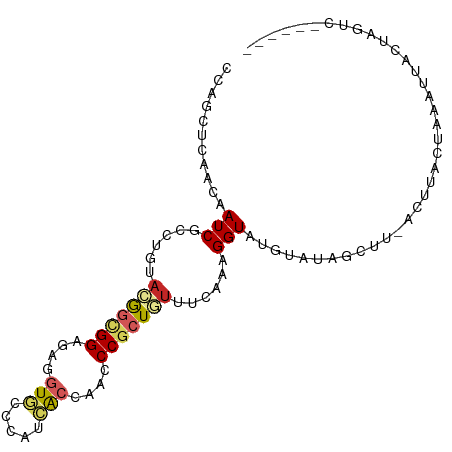
>dm3.chr3R 25679769 90 + 27905053 CCAGCUCAACAAUCGCCUGUACGGCGGAGAGGUGCCCAUCACCAACCCGCUGUUUCAAAGGUAUGCAUAGCUU-ACUUCCUAGUUUAUAAC-------- ..((((...((...((((..(((((((...((((.....))))...))))))).....)))).))...)))).-.................-------- ( -22.70, z-score = -1.21, R) >droMoj3.scaffold_6540 26933497 99 - 34148556 CCAGCAGAAUAAUCGCAUAUAUGCUGGCGAGCUACCCAUUGCCAAUCCGUUAUUUCAAAGGUAAAUUCUAGUCAGAUCUCAAAUAAAUGAUCAUCCACU ((((((..(((......))).)))))).((.(((....(((((................)))))....))))).((((..........))))....... ( -16.19, z-score = -0.55, R) >droVir3.scaffold_13047 2506345 98 + 19223366 CCAGCAAAAUAAUCGUUUAUACGCUGGCGAAUUGCCCAUAGCCAAUCCGUUGUUUCAAAGGUGGGUU-UAUUUAAUUUUCUCCCUCUUAAUUCUUAGCA (((((...((((....))))..))))).(((((....(((((......))))).....(((.(((..-.............))).))))))))...... ( -15.26, z-score = 0.29, R) >droWil1.scaffold_181089 2035595 92 + 12369635 CAAUCACAAUAAUCGCGUCUUUGGCGGUGAGGUUCCCAUUACGAAUCCACUUUUUCAACGGUAAGAGU--UUAACUCCACAAAAUUUAUAUACA----- ...........(((((((.....)))(((.(((((.......))))))))........))))..(((.--....))).................----- ( -12.60, z-score = 0.21, R) >droPer1.super_7 3861477 96 + 4445127 CCAACUGAACAAUCGCAUGUACAGCGGAGAGGUGCCCAUCACCAACCCGCUUUUCCAACGGUAGAUACAGCCAAAUAUAGCCAAAUAGUGCUCGUA--- ..............((..(((((((((...((((.....))))...)))))........(((.......)))...............))))..)).--- ( -19.20, z-score = -0.29, R) >dp4.chr2 1076182 96 + 30794189 CCAACUGAACAAUCGCAUGUACAGCGGAGAAGUGCCCAUCACCAACCCGCUUUUCCAACGGUAGAUACAGCAAAAUAUAGCCAAAUAGUGCUCGUA--- ((..(((.(((......))).))).(((((((((.............)))))))))...))....(((((((...(((......))).)))).)))--- ( -19.72, z-score = -1.19, R) >droEre2.scaffold_4820 8147455 92 - 10470090 CCAGCUCAACAAUCGCCUGUACGGCGGGGAGGUGCCCAUCACCAACCCGCUGUUUCAAAGGUAUGCAUACAUG-GCUUACUCAAUGUCCAGUU------ ..((((..(((...((((..((((((((..((((.....))))..)))))))).....))))..((.......-))........)))..))))------ ( -29.10, z-score = -1.52, R) >droYak2.chr3R 8045802 88 - 28832112 CCAGCUCAACAAUCGCCUGUACGGCGGAGAGGUGCCCAUCACCAAUCCGCUGUUCCAAAGGUAUAUAUA-----GCAUACGCAAUUCCAAGUU------ ...(((........((((..((((((((..((((.....))))..)))))))).....))))......)-----)).................------ ( -25.64, z-score = -2.90, R) >droSec1.super_4 4530059 92 + 6179234 CCAGCUCAACAAUCGCCUUUACGGCGGAGAGGUGCCCAUCACCAACCCACUGUUUCAAAGGUAUUCAUAGCUU-ACUUGCUAUUUUCCUAGUU------ .......(((....((((((((((.((...((((.....))))...)).))))...))))))....(((((..-....))))).......)))------ ( -20.50, z-score = -1.42, R) >droSim1.chr3R 25354477 92 + 27517382 CCAGCUCAACAAUCGCCUGUACGGCGGAGAGGUGCCCAUCACCAACCCGCUGUUUCAAAGGUAUGCACAGCUU-ACUUCCCAGUUUCCCAGUU------ ..((((...((...((((..(((((((...((((.....))))...))))))).....)))).))...)))).-...................------ ( -22.70, z-score = -0.94, R) >consensus CCAGCUCAACAAUCGCCUGUACGGCGGAGAGGUGCCCAUCACCAACCCGCUGUUUCAAAGGUAUGUAUAGCUU_ACUUACUAAAUUACUAGUC______ ...........(((......(((((((....(((.....)))....)))))))......)))..................................... ( -7.75 = -8.25 + 0.50)


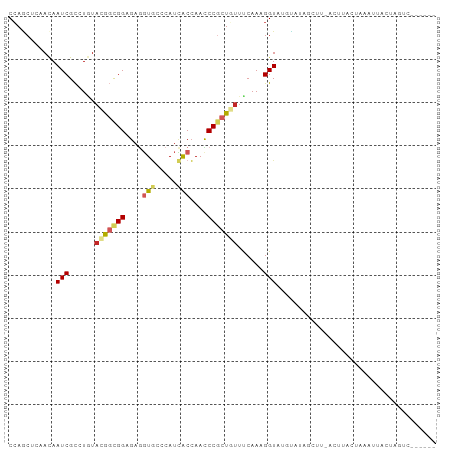
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:55:48 2011