
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 9,126,099 – 9,126,195 |
| Length | 96 |
| Max. P | 0.785235 |

| Location | 9,126,099 – 9,126,195 |
|---|---|
| Length | 96 |
| Sequences | 9 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 74.35 |
| Shannon entropy | 0.50593 |
| G+C content | 0.46003 |
| Mean single sequence MFE | -24.95 |
| Consensus MFE | -11.08 |
| Energy contribution | -11.62 |
| Covariance contribution | 0.54 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.68 |
| SVM RNA-class probability | 0.785235 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 9126099 96 + 23011544 CUCAACCGA-----CCCGGCCAAUAAACAGCUUC--AAAAGUGCCC-AAGAAUCGUUGCGAAUG--CGAAAAGAUACAUUUGUAUCUUUGGUCAUGCACUUUGCAG-- .....(((.-----..)))..........((...--.(((((((((-.....((((.......)--)))((((((((....))))))))))....)))))))))..-- ( -20.90, z-score = -0.96, R) >droSim1.chr2L 8906199 96 + 22036055 CUCAACCGA-----CCCGGCCAAUAAACAGCUUG--AAAAGUGCCC-AAGAAUCGUUGCGAAUG--CGAAAAGAUACAUUUGUAUCUUUGGCCAUGCACUUUGCAG-- .........-----...((((.........((((--.........)-)))..((((.......)--)))((((((((....)))))))))))).(((.....))).-- ( -22.90, z-score = -1.51, R) >droSec1.super_3 4592196 96 + 7220098 CUCAACCGA-----CCCGGCCAAUAAACAGCUUG--AUAAGUGCCC-AAGAAUCGUUGCGAAUG--CGAAAAGAUACAUUUGUAUCUUUGGCCAUGCACUUUGCAG-- .........-----...((((.........((((--.........)-)))..((((.......)--)))((((((((....)))))))))))).(((.....))).-- ( -22.90, z-score = -1.50, R) >droYak2.chr2L 11792641 98 + 22324452 CUCAUACGA---AACCCGGCCAAUAAACAGCUUG--AAAAGUGCCC-AAGAAUCGUUGCGAAUG--CGAAAAGAUACAUUUGUAUCUUUGGGCAUGCACUUUGCAG-- ......((.---....))...........((...--....((((((-.....((((.......)--)))((((((((....)))))))))))))).......))..-- ( -24.64, z-score = -1.59, R) >droEre2.scaffold_4929 9730561 101 + 26641161 CUCAUACGACCCGACCCGGCCAAUAAACAGCUUG--AAAAGUGCCC-AAGAAUCGUUGCGAAUG--CGAAAAGAUACAUUUGUAUCUUUGGGCAUGCACUUUGCAG-- ..........(((...)))..........((...--....((((((-.....((((.......)--)))((((((((....)))))))))))))).......))..-- ( -25.24, z-score = -1.59, R) >droAna3.scaffold_12943 474500 98 + 5039921 CAGAAACGC---CAAACAGCCAAUAAACGGCACG--AAAAGUGGCC-AAGAAUCGGUGCGAAUG--CCAAAAGAUACUUUUGUAUCUUUCGGCAUGCACUUUGCAG-- .......((---(.....(((.......)))((.--....))))).-.......(((((..(((--((.((((((((....)))))))).))))))))))......-- ( -30.90, z-score = -3.23, R) >dp4.chr4_group5 1432723 84 + 2436548 -------CA---GAGGCAGCCAAUAAACAGCUUCGCACAAGUGCCCCAAGAAUCGGUGCGAUUG--CCA---GAUACUUUUGUAUCUU-------GCACUUUGCAG-- -------..---(((((............)))))(((.((((((..........(((......)--))(---(((((....)))))).-------)))))))))..-- ( -25.20, z-score = -2.16, R) >droPer1.super_5 5839362 84 + 6813705 -------CA---GCGGCAGCCAAUAAACAGCUUCGCACAAGUGCCCCAAGAAUCGGUGCGAUUG--CCA---GAUACUUUUGUAUCUU-------GCACUUUGCAG-- -------..---((((.(((.........)))))))....................(((((.((--(.(---(((((....)))))).-------)))..))))).-- ( -22.60, z-score = -0.93, R) >droVir3.scaffold_12963 476266 99 + 20206255 --CAAACCU-----AAGAAUUAUGCACAAGCACA--AGCACAGCGCCAGAAAUUGGCUCGAACGUACUCAAAGAUACAUUUGUAUCUUUUGCCUUGCAGUUUGUGCAG --.......-----........(((((((((...--.(((..(((((((...)))))............((((((((....)))))))).))..))).))))))))). ( -29.30, z-score = -3.12, R) >consensus CUCAAACGA_____CCCGGCCAAUAAACAGCUUG__AAAAGUGCCC_AAGAAUCGUUGCGAAUG__CGAAAAGAUACAUUUGUAUCUUUGGCCAUGCACUUUGCAG__ ......................................((((((........(((...)))........((((((((....))))))))......))))))....... (-11.08 = -11.62 + 0.54)


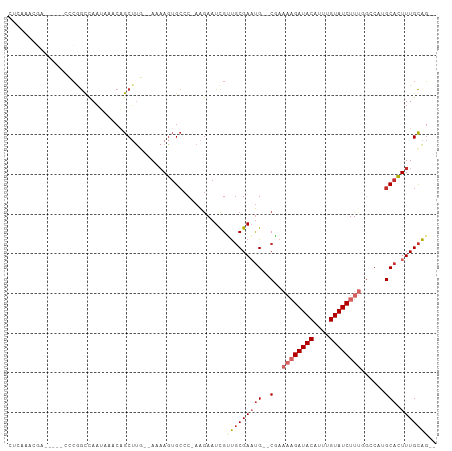
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:27:38 2011