
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 25,540,940 – 25,541,070 |
| Length | 130 |
| Max. P | 0.563877 |

| Location | 25,540,940 – 25,541,032 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 93 |
| Reading direction | reverse |
| Mean pairwise identity | 81.44 |
| Shannon entropy | 0.30005 |
| G+C content | 0.47437 |
| Mean single sequence MFE | -29.65 |
| Consensus MFE | -20.16 |
| Energy contribution | -21.85 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.64 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.14 |
| SVM RNA-class probability | 0.563877 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 25540940 92 - 27905053 UCCCUCCUGGGAUUGGCCUUAUCCCAGGAUCGAAAUGAUACGGAGUGGAUCCAAAGUCAGAAGGAUCGUGAGUAGUUUAGGCUGCCUUUUCA- ((..(((((((((.......)))))))))..))........((((.((((((..........)))))....(((((....)))))).)))).- ( -27.00, z-score = -0.17, R) >droYak2.chr3R 7903545 93 + 28832112 UCCCUCCUGGGAAUGGCUGUCUCCCAGAAUAGAACUGAAGCGGAAAGGCAGAAAACUCAGGAGGAAUGUAAGUUGUUUAAACUUCCUUUUUUA ..(((((((((.....(((((((((((.......)))....)))..)))))....)))))))))...(.(((((.....))))).)....... ( -26.90, z-score = -1.07, R) >droSec1.super_4 4395051 92 - 6179234 UCCCUCCUGGGAUUGGCCGUAUCCCAGGAUCGAAAUAGUACGGAGUGGAUGAAAAGUCAGAAGGAUCGUGAGUAGUUUAAGCUGUCUUUUCA- (((((((((((((.......)))))))))(((........))).).)))(((((((.(((...((.(....)....))...))).)))))))- ( -29.00, z-score = -1.87, R) >droSim1.chr3R 25213518 92 - 27517382 UCCCUCCUGGGAUUGGCCGUAUCCCAGGACCGAAAUAGUCCGGAGUGGAAGAAAAGUCAGAUGGAGCGUGAGUAGUUUAAGCUGCCUUUUCA- (((((((((((((.......))))))((((.......)))))))).))).((((((.(((...((((.......))))...))).)))))).- ( -35.70, z-score = -3.44, R) >consensus UCCCUCCUGGGAUUGGCCGUAUCCCAGGAUCGAAAUAAUACGGAGUGGAAGAAAAGUCAGAAGGAUCGUGAGUAGUUUAAGCUGCCUUUUCA_ ..(((((((((((.......)))))))))(((........)))...))..((((((.(((...((((.......))))...))).)))))).. (-20.16 = -21.85 + 1.69)



| Location | 25,540,965 – 25,541,070 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 67.21 |
| Shannon entropy | 0.55791 |
| G+C content | 0.50286 |
| Mean single sequence MFE | -32.74 |
| Consensus MFE | -15.91 |
| Energy contribution | -17.07 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.547109 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 25540965 105 + 27905053 ------------GAUCCUUCUGACUUUGGAUCCACUCCGUAUCAUUUCGAUCCUGGGAUAAGGCCAAUCCCAGGAGGGAAGCCAAUACGACCAAGAGGAGAAAUAGCGGAGUCUUCA ------------(((((..........))))).(((((((...(((((..(((((((((.......)))))))))((....)).......((....)).))))).)))))))..... ( -36.90, z-score = -2.01, R) >droEre2.scaffold_4784 12459336 105 - 25762168 UUGGAAUUGGAAAUUUGACCGGAGGUUUCUUUUGCCACGGCCAAGUCCAAGACUGGA------CCAAACCGACUAUGCAAACGAG---AACCAGGAGAAGUGAUAC---AGCUUUCA .((((((((...((((..((...((((((.(((((..(((....(((((....))))------)....))).....))))).)))---.))).))..))))....)---)).))))) ( -24.50, z-score = -0.19, R) >droYak2.chr3R 7903571 105 - 28832112 ------------AUUCCUCCUGAGUUUUCUGCCUUUCCGCUUCAGUUCUAUUCUGGGAGACAGCCAUUCCCAGGAGGGAGGCCAAUAUGACCAAGAGGAGGAACACCUGGGUCUUCA ------------.((((((.((.(((....((((..(.((....))....(((((((((.......))))))))))..))))......))))).))))))(((.((....)).))). ( -33.00, z-score = 0.30, R) >droSec1.super_4 4395076 105 + 6179234 ------------GAUCCUUCUGACUUUUCAUCCACUCCGUACUAUUUCGAUCCUGGGAUACGGCCAAUCCCAGGAGGGAAGCCAAUACGACCAAGAGGAGAAACAGCGGAGUCUUCA ------------((...(((((.((((((.(((.((.((((...((((..(((((((((.......)))))))))..))))....))))....)).))))))).)))))))...)). ( -32.40, z-score = -1.53, R) >droSim1.chr3R 25213543 105 + 27517382 ------------GCUCCAUCUGACUUUUCUUCCACUCCGGACUAUUUCGGUCCUGGGAUACGGCCAAUCCCAGGAGGGAAGCCAAUACGACCAAGAGGAGAAACAGCGGAGUCUUCA ------------(((((..(((......(((((.(((((((((.....)))))((((((.......))))))))))))))).........((....)).....))).)))))..... ( -36.90, z-score = -1.93, R) >consensus ____________GAUCCUUCUGACUUUUCUUCCACUCCGGACCAUUUCGAUCCUGGGAUACGGCCAAUCCCAGGAGGGAAGCCAAUACGACCAAGAGGAGAAACAGCGGAGUCUUCA .....................(((((........((((............(((((((((.......)))))))))((....)).............))))........))))).... (-15.91 = -17.07 + 1.16)


| Location | 25,540,965 – 25,541,070 |
|---|---|
| Length | 105 |
| Sequences | 5 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 67.21 |
| Shannon entropy | 0.55791 |
| G+C content | 0.50286 |
| Mean single sequence MFE | -34.02 |
| Consensus MFE | -16.04 |
| Energy contribution | -19.28 |
| Covariance contribution | 3.24 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.13 |
| Structure conservation index | 0.47 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.513952 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 25540965 105 - 27905053 UGAAGACUCCGCUAUUUCUCCUCUUGGUCGUAUUGGCUUCCCUCCUGGGAUUGGCCUUAUCCCAGGAUCGAAAUGAUACGGAGUGGAUCCAAAGUCAGAAGGAUC------------ .....((((((.((((((...((((((..(((..((((((((....))))..)))).))).))))))..))))))...)))))).(((((..........)))))------------ ( -36.00, z-score = -1.38, R) >droEre2.scaffold_4784 12459336 105 + 25762168 UGAAAGCU---GUAUCACUUCUCCUGGUU---CUCGUUUGCAUAGUCGGUUUGG------UCCAGUCUUGGACUUGGCCGUGGCAAAAGAAACCUCCGGUCAAAUUUCCAAUUCCAA .((((..(---(..........((.((((---.((.(((((.....((((..((------(((......)))))..))))..))))).))))))...)).))..))))......... ( -26.70, z-score = -0.93, R) >droYak2.chr3R 7903571 105 + 28832112 UGAAGACCCAGGUGUUCCUCCUCUUGGUCAUAUUGGCCUCCCUCCUGGGAAUGGCUGUCUCCCAGAAUAGAACUGAAGCGGAAAGGCAGAAAACUCAGGAGGAAU------------ .(((.((....)).)))........((((.....))))..(((((((((.....(((((((((((.......)))....)))..)))))....)))))))))...------------ ( -33.60, z-score = -0.26, R) >droSec1.super_4 4395076 105 - 6179234 UGAAGACUCCGCUGUUUCUCCUCUUGGUCGUAUUGGCUUCCCUCCUGGGAUUGGCCGUAUCCCAGGAUCGAAAUAGUACGGAGUGGAUGAAAAGUCAGAAGGAUC------------ .....(((((((((((((...((((((..((((.((((((((....))))..)))))))).))))))..)))))))..)))))).(((.....))).........------------ ( -36.50, z-score = -1.63, R) >droSim1.chr3R 25213543 105 - 27517382 UGAAGACUCCGCUGUUUCUCCUCUUGGUCGUAUUGGCUUCCCUCCUGGGAUUGGCCGUAUCCCAGGACCGAAAUAGUCCGGAGUGGAAGAAAAGUCAGAUGGAGC------------ ......(((((((((((((((((((((..((((.((((((((....))))..)))))))).))))))(((........)))...)).)))))...))).))))).------------ ( -37.30, z-score = -1.44, R) >consensus UGAAGACUCCGCUGUUUCUCCUCUUGGUCGUAUUGGCUUCCCUCCUGGGAUUGGCCGUAUCCCAGGAUCGAAAUAGUACGGAGUGGAAGAAAAGUCAGAAGGAAC____________ .....((((((.((((((...((((((......(((((((((....))))..)))))....))))))..))))))...))))))................................. (-16.04 = -19.28 + 3.24)
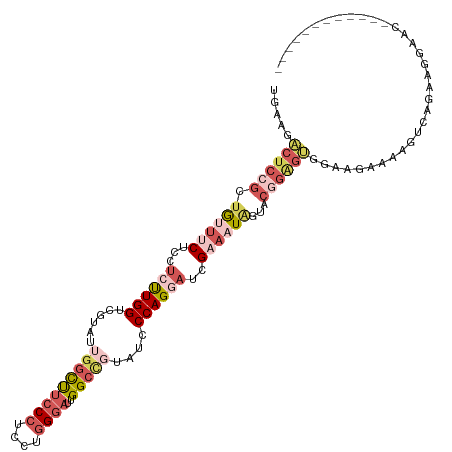


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:55:21 2011