| Sequence ID | dm3.chr3R |
|---|---|
| Location | 25,293,292 – 25,293,364 |
| Length | 72 |
| Max. P | 0.939118 |

| Location | 25,293,292 – 25,293,364 |
|---|---|
| Length | 72 |
| Sequences | 4 |
| Columns | 72 |
| Reading direction | forward |
| Mean pairwise identity | 56.44 |
| Shannon entropy | 0.72287 |
| G+C content | 0.29797 |
| Mean single sequence MFE | -11.71 |
| Consensus MFE | -4.28 |
| Energy contribution | -4.52 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.37 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.45 |
| SVM RNA-class probability | 0.939118 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
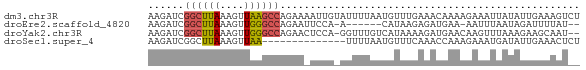
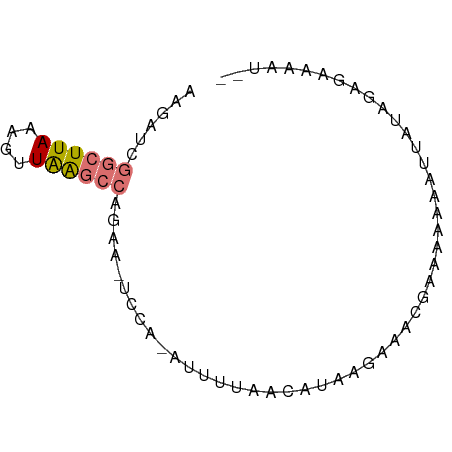
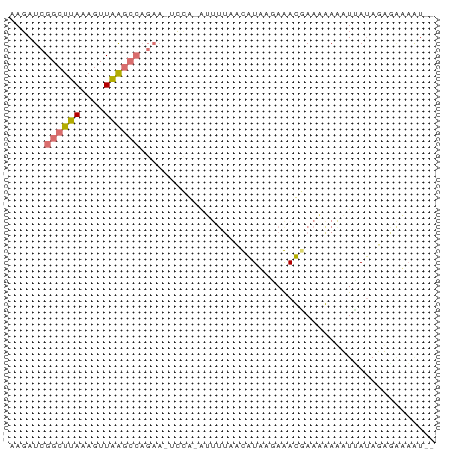
>dm3.chr3R 25293292 72 + 27905053 AAGAUCGGCUUAAAGUUAAGCCAGAAAAUUGUAUUUUAAUGUUUGAAACAAAAGAAAUUAUAUUGAAAGUCU .((((.((((((....))))))...........(((((((((.................))))))))))))) ( -11.53, z-score = -1.64, R) >droEre2.scaffold_4820 7757044 62 - 10470090 AAGAUCGGCUUAAAGUUGGGCCAGAAUUCCA-A------CAUAAGAGAUGAA-AAUUUAAUAGAUUUUAU-- ((((((((((((....)))))).........-.------......((((...-.))))....))))))..-- ( -11.70, z-score = -1.95, R) >droYak2.chr3R 7640618 69 - 28832112 AAGAUCGGCUUAAAGUUGGGCCAGAACUCCA-GGUUUGUCAUAAAAGAUGAACAAGUUUAAAGAAGCAAU-- ....((((((((....)))))).(((((...-.(((..((......))..))).)))))...))......-- ( -15.50, z-score = -1.90, R) >droSec1.super_4 4152141 58 + 6179234 AAGAUCGGCUUAAAGUUAA--------------UUUUAAUGUUUCAAACCAAAGAAAUGAUAUUGAAACUCU (((((..((.....))..)--------------))))...(((((((.....(....)....)))))))... ( -8.10, z-score = -1.15, R) >consensus AAGAUCGGCUUAAAGUUAAGCCAGAA_UCCA_AUUUUAACAUAAGAAACGAAAAAAAUUAUAGAGAAAAU__ ......((((((....)))))).................................................. ( -4.28 = -4.52 + 0.25)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:54:35 2011