
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 25,279,419 – 25,279,525 |
| Length | 106 |
| Max. P | 0.740914 |

| Location | 25,279,419 – 25,279,525 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 81.24 |
| Shannon entropy | 0.32406 |
| G+C content | 0.42354 |
| Mean single sequence MFE | -31.44 |
| Consensus MFE | -17.00 |
| Energy contribution | -17.48 |
| Covariance contribution | 0.48 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.25 |
| SVM RNA-class probability | 0.615022 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
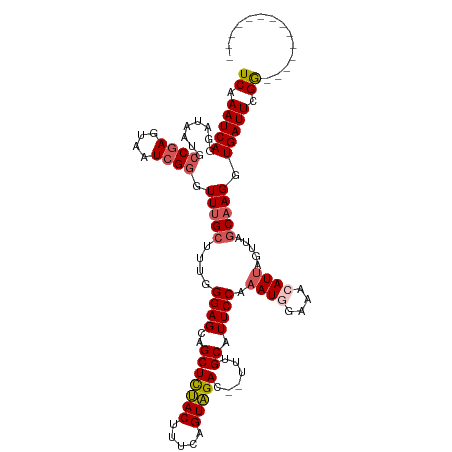
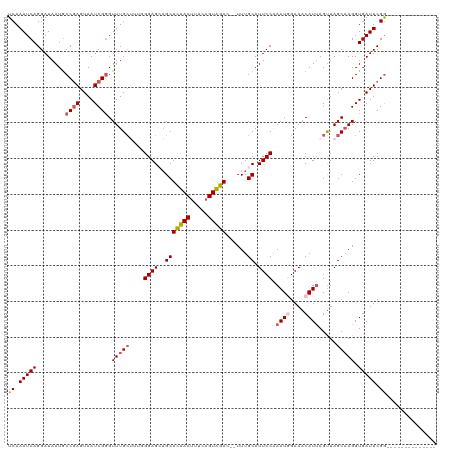
>dm3.chr3R 25279419 106 + 27905053 UCAAAUCAGGAUAAUACCGAGUAAUCGGGUUUGCUUUGGGAGAAGCUCUACUUUCAGUAGAGCAUUUGCAUUCCAAAUGGAAACAUUAGCUAGCCAGGUGAUUCGG-------------- ................((((((.(((.((((.(((.(((((((((((((((.....))))))).)))...)))))((((....))))))).)))).))).))))))-------------- ( -36.40, z-score = -4.06, R) >droEre2.scaffold_4820 7740999 118 - 10470090 UCAAAUCAGGAUAAUGCCGAGUAAUCGGGUUUGCGAUUGGAGCUGCUUUACUACCAGUAGAG--UUUGCAUUCCGAAUUGAAACAUCAGUUAGCAAGGUGAUUCGGGAUGAUUCCCUCGC ...(((((........((((....)))).(((((...((((((.(((((((.....))))))--)..))..))))((((((....)))))).))))).))))).((((....)))).... ( -34.20, z-score = -1.42, R) >droYak2.chr3R 7626231 118 - 28832112 UCAAAUCAGGAUAAUGCCAAGUAAUCGGGUUUGCUCAUGGAGAAGCUUCACUUUCUGUGGAA--UUUGCAUUCCAAAUUGAAACAUUAGUUAGCAAGGUGAUUCGAGAGGGAUGCCUUAC .((((((..(((.((.....)).))).)))))).(((((((((........)))))))))..--...(((((((...(((((.((((.........)))).)))))..)))))))..... ( -25.90, z-score = 0.43, R) >droSec1.super_4 4138115 99 + 6179234 UCAAAUCAGGAUAAUGCCGAGUAAUCGUGUUUG-----GGAGCAGCUCUACUUUCAGUAGAC--UUUGCAUUCCAAAUGGAAACAUUAGUUAGCAAGGUGAUUCGG-------------- ................((((((.(((.((((((-----(((((((.(((((.....))))).--.)))).)))))((((....))))....)))).))).))))))-------------- ( -28.70, z-score = -2.66, R) >droSim1.chr3R 24952281 104 + 27517382 UCAAAUCAGGAUAAUGCCGAGUAAUCGGGUUUGCUUUGGGAGCAGCUCUACUUUCAGUAGAC--UUUGCAUUCCAAAUGGAAACAUUAGUUAGCAAGGUGAUUCGG-------------- .(.(((((........((((....)))).((((((.(((((((((.(((((.....))))).--.)))).)))))((((....))))....)))))).))))).).-------------- ( -32.00, z-score = -2.86, R) >consensus UCAAAUCAGGAUAAUGCCGAGUAAUCGGGUUUGCUUUGGGAGCAGCUCUACUUUCAGUAGAC__UUUGCAUUCCAAAUGGAAACAUUAGUUAGCAAGGUGAUUCGG______________ ................((((((.(((..(((.((....((((..(((((((.....)))))......)).)))).((((....)))).)).)))..))).)))))).............. (-17.00 = -17.48 + 0.48)

| Location | 25,279,419 – 25,279,525 |
|---|---|
| Length | 106 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.24 |
| Shannon entropy | 0.32406 |
| G+C content | 0.42354 |
| Mean single sequence MFE | -25.36 |
| Consensus MFE | -16.34 |
| Energy contribution | -16.50 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.88 |
| Structure conservation index | 0.64 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.740914 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
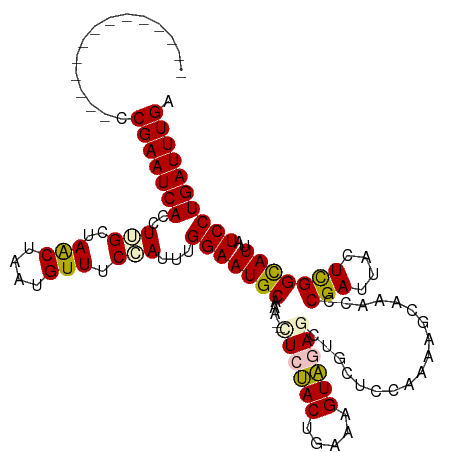
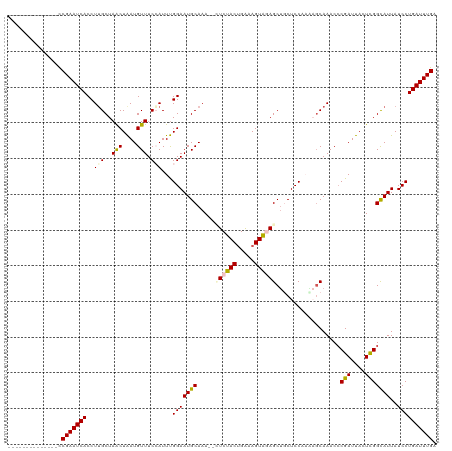
>dm3.chr3R 25279419 106 - 27905053 --------------CCGAAUCACCUGGCUAGCUAAUGUUUCCAUUUGGAAUGCAAAUGCUCUACUGAAAGUAGAGCUUCUCCCAAAGCAAACCCGAUUACUCGGUAUUAUCCUGAUUUGA --------------.(((((((...((.(((((..(((((((....)))).)))...(((((((.....))))))).........)))....((((....))))...)).))))))))). ( -28.90, z-score = -3.22, R) >droEre2.scaffold_4820 7740999 118 + 10470090 GCGAGGGAAUCAUCCCGAAUCACCUUGCUAACUGAUGUUUCAAUUCGGAAUGCAAA--CUCUACUGGUAGUAAAGCAGCUCCAAUCGCAAACCCGAUUACUCGGCAUUAUCCUGAUUUGA ((((((((.((.....)).)).))))))...........((((.(((((((((...--.......((.(((......)))))(((((......))))).....)))))...)))).)))) ( -27.80, z-score = -0.97, R) >droYak2.chr3R 7626231 118 + 28832112 GUAAGGCAUCCCUCUCGAAUCACCUUGCUAACUAAUGUUUCAAUUUGGAAUGCAAA--UUCCACAGAAAGUGAAGCUUCUCCAUGAGCAAACCCGAUUACUUGGCAUUAUCCUGAUUUGA ...(((....))).((((((((...((((((.((((..((((...((((((....)--))))).......))))((((......)))).......)))).))))))......)))))))) ( -24.10, z-score = -0.77, R) >droSec1.super_4 4138115 99 - 6179234 --------------CCGAAUCACCUUGCUAACUAAUGUUUCCAUUUGGAAUGCAAA--GUCUACUGAAAGUAGAGCUGCUCC-----CAAACACGAUUACUCGGCAUUAUCCUGAUUUGA --------------.(((((((...(((...............(((((...(((..--.(((((.....)))))..)))..)-----))))..(((....))))))......))))))). ( -21.20, z-score = -1.78, R) >droSim1.chr3R 24952281 104 - 27517382 --------------CCGAAUCACCUUGCUAACUAAUGUUUCCAUUUGGAAUGCAAA--GUCUACUGAAAGUAGAGCUGCUCCCAAAGCAAACCCGAUUACUCGGCAUUAUCCUGAUUUGA --------------.(((((((..((((((((....))).....((((...(((..--.(((((.....)))))..)))..)))))))))..((((....))))........))))))). ( -24.80, z-score = -2.64, R) >consensus ______________CCGAAUCACCUUGCUAACUAAUGUUUCCAUUUGGAAUGCAAA__CUCUACUGAAAGUAGAGCUGCUCCAAAAGCAAACCCGAUUACUCGGCAUUAUCCUGAUUUGA ...............(((((((..(((..(((....))).)))...(((((((.....((((((.....))))))..................(((....)))))))..)))))))))). (-16.34 = -16.50 + 0.16)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:54:33 2011