
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 25,264,707 – 25,264,805 |
| Length | 98 |
| Max. P | 0.797325 |

| Location | 25,264,707 – 25,264,805 |
|---|---|
| Length | 98 |
| Sequences | 9 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 64.90 |
| Shannon entropy | 0.71141 |
| G+C content | 0.56132 |
| Mean single sequence MFE | -20.01 |
| Consensus MFE | -6.98 |
| Energy contribution | -7.62 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.31 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.797325 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 25264707 98 + 27905053 -GCUG-CCGCAGACAAAACUAUGCACAUUUAACGCUAAAUAUGCCAACUGCCACUCCCCCUUUU-CAGCAAGGACCACUCCCCUGGCCGCCGUCCCCUUUU-- -((.(-((((((..........(((.(((((....))))).)))...)))).......((((..-....))))...........))).))...........-- ( -15.82, z-score = -0.45, R) >droAna3.scaffold_12911 4953763 90 + 5364042 --CCCGCCGCAUACAAAACUGAGCCCAUUUAGCGCCAAAUGGGCUGUG-AUUGCCCCUCCUCCUGGA------CUUGAGU--CCACCCUCUACUUAGGCCU-- --...((((((......((..(((((((((......))))))))))).-..))).........((((------(....))--)))...........)))..-- ( -25.30, z-score = -2.08, R) >droEre2.scaffold_4820 7726308 80 - 10470090 -GCUG-CCGCAGACAAAACUAUGCACAUUUAACGCUAAAUGUGCCGGCUGCCACUCCUCCUUU-----------CCA--------GCCUCCGCCCCCUUUU-- -....-..((.((.........(((((((((....))))))))).(((((.............-----------.))--------))))).))........-- ( -17.94, z-score = -2.60, R) >droYak2.chr3R 7611134 82 - 28832112 -GCUG-CCGCAGACAAAACUAUGCACAUUUAACGCUAAAUGUGCCGGCUGCCACUCCCCC------------AACCACCA-----GCCUCCGCCCCCUUUU-- -....-..((.((.........(((((((((....))))))))).(((((..........------------......))-----))))).))........-- ( -17.79, z-score = -2.74, R) >droSec1.super_4 4123604 97 + 6179234 -GCUG-CCGCAGACAAAACUAUGCACAUUUAACGCUAAAUGUGCCGCCUGCCACUCCCCCUUUUUCAGCAAGGACCACUC--CUGGCCGCCGUCCCCUUUU-- -((.(-((((((..........(((((((((....)))))))))...))))...................((((....))--))))).))...........-- ( -22.32, z-score = -1.84, R) >droSim1.chr3R 24940210 96 + 27517382 -GCUG-CCGCAGACAAAACUAUGCACAUUUAACGCUAAAUGUGCCGCCUGCCACUCCCCCUUCU-CAGCAAGGACCACUC--CUGGCCGCCGCCCCCUUUU-- -((.(-((((((..........(((((((((....)))))))))...)))).............-.....((((....))--))))).))...........-- ( -22.32, z-score = -1.89, R) >droPer1.super_7 2286473 95 + 4445127 --CCUGCCGCAGACAAAACUAUGCACAUUUAACGCUAAAUGGGUUGUCUGCCACUCCCACUCCCACU------UCCCAGUUGCCAGCCUCUUCUUUGGUGCCU --......((((((((..(...((.........)).....)..))))))))................------....((..(((((........)))))..)) ( -18.60, z-score = -0.18, R) >dp4.chr2 2118696 101 + 30794189 --CCUGCCGCAGACAAAACUAUGCACAUUUAACGCUAAAUGGGUUGUCUGCCACUCCCACUCCCACUCCUACUUCCCAGUUGCCAGCCUCUUCCUUGGUGCCU --......((((((((..(...((.........)).....)..))))))))..........................((..(((((........)))))..)) ( -18.60, z-score = -0.34, R) >droVir3.scaffold_12822 2622534 79 + 4096053 UGCUGGCCCCAGACAAAACUCUGCAAAUUUAACGCCAAAUGGGCGUGCCGCAAUCCGCACAU-----------GCCCCCC-----GACGCCGCCC-------- .((.(((..((((......)))).................(((((((..((.....)).)))-----------))))...-----...)))))..-------- ( -21.40, z-score = -1.04, R) >consensus _GCUG_CCGCAGACAAAACUAUGCACAUUUAACGCUAAAUGUGCCGGCUGCCACUCCCCCUUCU_CA______ACCACGC__C__GCCUCCGCCCCCUUUU__ ........((((..........(((((((((....)))))))))...)))).................................................... ( -6.98 = -7.62 + 0.64)


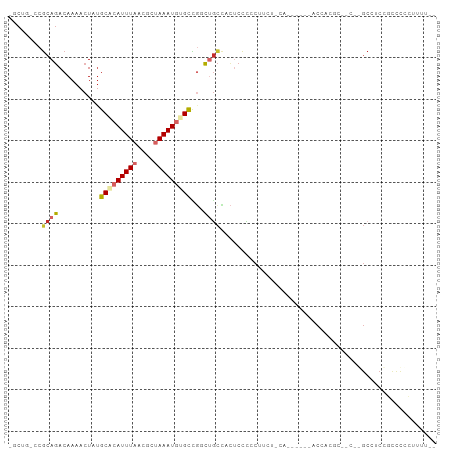
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:54:30 2011