
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 9,050,382 – 9,050,475 |
| Length | 93 |
| Max. P | 0.823685 |

| Location | 9,050,382 – 9,050,475 |
|---|---|
| Length | 93 |
| Sequences | 9 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 70.37 |
| Shannon entropy | 0.60499 |
| G+C content | 0.36074 |
| Mean single sequence MFE | -13.40 |
| Consensus MFE | -7.77 |
| Energy contribution | -8.08 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.25 |
| Mean z-score | -0.82 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.823685 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
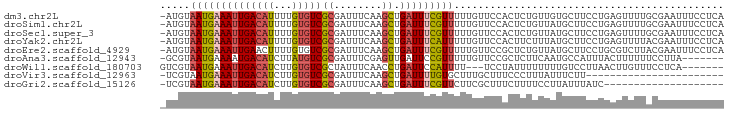
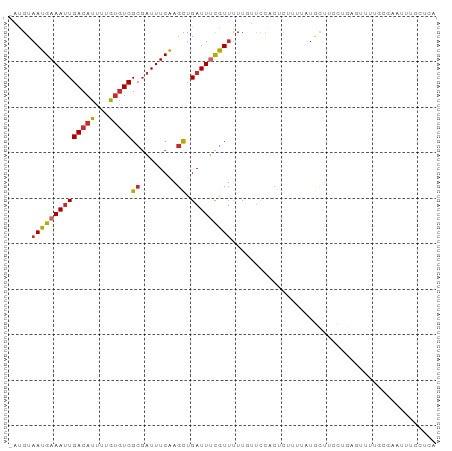
>dm3.chr2L 9050382 93 + 23011544 -AUGUAAUGAAAUUGACAUUUUGUGUCGCGAUUUCAAGCUGAUUUCGUUUUUGUUCCACUCUGUUGUGCUUCCUGAGUUUUGCGAAUUUCCUCA -....((((((((((((((...)))))((........)).)))))))))(((((...((((.............))))...)))))........ ( -16.62, z-score = -0.68, R) >droSim1.chr2L 8830836 93 + 22036055 -AUGUAAUGAAAUUGACAUUUUGUGUCGCGAUUUCAAGCUGAUUUCGUUUUUGUUCCACUCUGUUAUGCUUCCUGAGUUUUGCGAAUUUCCUCA -....((((((((((((((...)))))((........)).)))))))))(((((...((((.............))))...)))))........ ( -16.62, z-score = -0.92, R) >droSec1.super_3 4516688 93 + 7220098 -AUGUAAUGAAAUUGACAUUUUGUGUCGCGAUUUCAAGCUGAUUUCGUUUUUGUUCCACUCUGUUAUGCUUCCUGAGUUUUGCGAAUUUCCUCA -....((((((((((((((...)))))((........)).)))))))))(((((...((((.............))))...)))))........ ( -16.62, z-score = -0.92, R) >droYak2.chr2L 11710429 93 + 22324452 -AUGUAAUGAAAUUGACAUUUUGUGUCGCGAUUUCAAGCUGAUUUCAUUUUUGUUCCACUUCUUUAUGCUUCCUGAGUUUUACGAAUUUCCUCA -....((((((((((((((...)))))((........)).)))))))))(((((...((((.............))))...)))))........ ( -13.62, z-score = -0.54, R) >droEre2.scaffold_4929 9651710 93 + 26641161 -AUGUAAUGAAAUUGAACUUUUGUGUCGCGAUUUCAAGCUGAUUUCGUUUUUGUUCCGCUCUGUUAUGCUUCCUGCGUCUUACGAAUUUCCUCA -.(((((((((((((.((....))....))))))))(((.((...((....)).)).))).....((((.....)))).))))).......... ( -13.80, z-score = -0.16, R) >droAna3.scaffold_12943 390277 86 + 5039921 -GCGUAAUGAAAAUGACAUCUUAUGUCGCGAUUUCGAGUUGAUUCCGUUUUUGUUCCGCUCUUCAAUGCCAUUUACUUUUUUCCUUA------- -(((.((..((((((..(((.....(((......)))...)))..))))))..)).)))............................------- ( -13.10, z-score = -0.78, R) >droWil1.scaffold_180703 3848438 84 + 3946847 GUCGUAAUGAAAUUGACAUCUUGUGUCGCUAUUUCAACCUGAUUCCAUUUU---UCCUAUUUUUUUUGUCCUUAACUUGUUUCCUCA------- ((((...((((((((((((...))))))..))))))...))))........---.................................------- ( -8.30, z-score = -1.38, R) >droVir3.scaffold_12963 371290 70 + 20206255 -UCGUAAUGAAAUUGACAUCUUGUGUCGCGAUUUCAAGCUGAUUUUGUGCUUUGCUUUCCCUUUAUUUCUU----------------------- -..((((((((((((((((...)))))..)))))))(((.........)))))))................----------------------- ( -10.50, z-score = -0.88, R) >droGri2.scaffold_15126 5139789 73 - 8399593 -UCGUAAUGAAAUUGACAUCUUGUGUCGCGAUUUCAAGCUGAUUUCGUUCUUCGCUUUCUUUUCCUUAUUUAUC-------------------- -.((.((((((((((((((...)))))((........)).)))))))))...))....................-------------------- ( -11.40, z-score = -1.12, R) >consensus _AUGUAAUGAAAUUGACAUUUUGUGUCGCGAUUUCAAGCUGAUUUCGUUUUUGUUCCACUCUUUUAUGCUUCCUGAGUUUUGCGAAUUUCCUCA .....((((((((((((((...)))))((........)).)))))))))............................................. ( -7.77 = -8.08 + 0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:27:29 2011