
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 24,921,290 – 24,921,381 |
| Length | 91 |
| Max. P | 0.998693 |

| Location | 24,921,290 – 24,921,381 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 86.75 |
| Shannon entropy | 0.25297 |
| G+C content | 0.48164 |
| Mean single sequence MFE | -17.22 |
| Consensus MFE | -11.69 |
| Energy contribution | -12.38 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.11 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.64 |
| SVM RNA-class probability | 0.770996 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
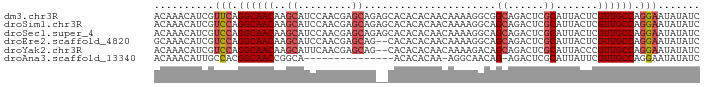
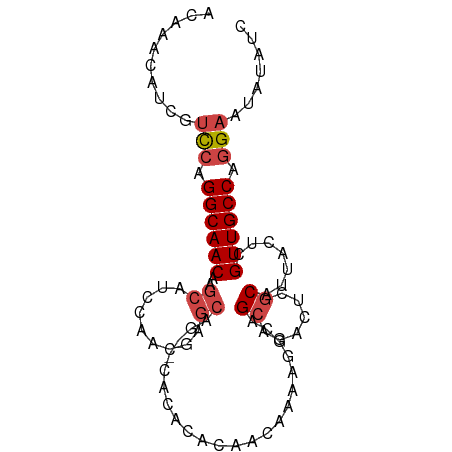
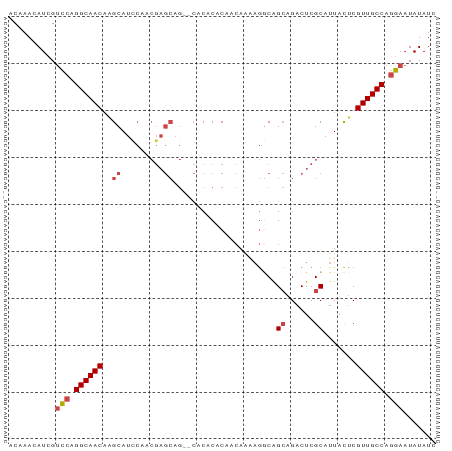
>dm3.chr3R 24921290 91 + 27905053 ACAAACAUCGUUCAGGCAACAAGCAUCCAACGAGCAGAGCACACACAACAAAAGGCGGCAGACUCGCAUUACUCGUUGCCAGGAAUAUAUC .........((((.((((((..((.((....)))).(((.........(....)((((.....))))....)))))))))..))))..... ( -18.20, z-score = -1.89, R) >droSim1.chr3R 24604632 91 + 27517382 ACAAACAUCGUCCAGGCAACAAGCAUCCAACGAGCAGAGCACACACAACAAAAGGCAGCAGACUCGCAUUACUCGUUGCCAGGAAUAUAUC ..........(((.((((((..((.((....)))).(((.........(....)...((......))....))))))))).)))....... ( -17.50, z-score = -1.99, R) >droSec1.super_4 3786026 91 + 6179234 ACAAACAUCGUCCAGGCAACAAGCAUCCAACGAGCAGAGCACACACAACAAAAGGCAGCAGACUCGCAUUACUCGUUGCCAGGAAUAUAUC ..........(((.((((((..((.((....)))).(((.........(....)...((......))....))))))))).)))....... ( -17.50, z-score = -1.99, R) >droEre2.scaffold_4820 7359744 89 - 10470090 GCAAACAUCGUCCAGGCAACAAGCAUCCAACGAGCAG--CACACACAACAAAAGGCAGCAGACUCGCAUUACUCGUUGCCAGGAAUAUAUC ..........(((.((((((.((.......(((((.(--(.................)).).)))).....)).)))))).)))....... ( -17.53, z-score = -1.84, R) >droYak2.chr3R 7255169 89 - 28832112 ACAAACAUCGUCCAGGCAACAAGCAUUCAACGAGCAG--CACACACAACAAAAGACAGCAGACUCGCAUUACCCGUUGCCAGGAAUAUAUC ..........(((.((((((..((.........))..--..................((......)).......)))))).)))....... ( -16.20, z-score = -2.56, R) >droAna3.scaffold_13340 4043079 74 - 23697760 ACAAACAUUGCCACGGCAACCGGCA---------------ACACACAA-AGGCAACAG-AGACUCGCAUUAUUCGUUGCCAGGAAUAUAUC .......((((....))))((((((---------------((......-.(....).(-.....).........)))))).))........ ( -16.40, z-score = -2.39, R) >consensus ACAAACAUCGUCCAGGCAACAAGCAUCCAACGAGCAG__CACACACAACAAAAGGCAGCAGACUCGCAUUACUCGUUGCCAGGAAUAUAUC ..........(((.((((((..((.........))......................((......)).......)))))).)))....... (-11.69 = -12.38 + 0.69)
| Location | 24,921,290 – 24,921,381 |
|---|---|
| Length | 91 |
| Sequences | 6 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 86.75 |
| Shannon entropy | 0.25297 |
| G+C content | 0.48164 |
| Mean single sequence MFE | -30.58 |
| Consensus MFE | -21.33 |
| Energy contribution | -22.92 |
| Covariance contribution | 1.59 |
| Combinations/Pair | 1.08 |
| Mean z-score | -4.01 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.45 |
| SVM RNA-class probability | 0.998693 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
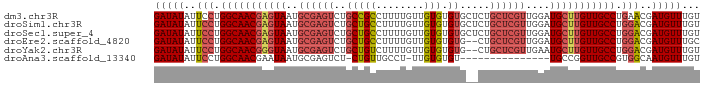
>dm3.chr3R 24921290 91 - 27905053 GAUAUAUUCCUGGCAACGAGUAAUGCGAGUCUGCCGCCUUUUGUUGUGUGUGCUCUGCUCGUUGGAUGCUUGUUGCCUGAACGAUGUUUGU (((((.(((..(((((((((((..((((((..((((((.......).))).))...))))))....))))))))))).)))..)))))... ( -31.80, z-score = -4.36, R) >droSim1.chr3R 24604632 91 - 27517382 GAUAUAUUCCUGGCAACGAGUAAUGCGAGUCUGCUGCCUUUUGUUGUGUGUGCUCUGCUCGUUGGAUGCUUGUUGCCUGGACGAUGUUUGU (((((..(((.(((((((((((..((((((..((((((.......).))).))...))))))....))))))))))).)))..)))))... ( -32.90, z-score = -4.49, R) >droSec1.super_4 3786026 91 - 6179234 GAUAUAUUCCUGGCAACGAGUAAUGCGAGUCUGCUGCCUUUUGUUGUGUGUGCUCUGCUCGUUGGAUGCUUGUUGCCUGGACGAUGUUUGU (((((..(((.(((((((((((..((((((..((((((.......).))).))...))))))....))))))))))).)))..)))))... ( -32.90, z-score = -4.49, R) >droEre2.scaffold_4820 7359744 89 + 10470090 GAUAUAUUCCUGGCAACGAGUAAUGCGAGUCUGCUGCCUUUUGUUGUGUGUG--CUGCUCGUUGGAUGCUUGUUGCCUGGACGAUGUUUGC (((((..(((.(((((((((((..((((((..((((((.......).))).)--).))))))....))))))))))).)))..)))))... ( -33.10, z-score = -4.53, R) >droYak2.chr3R 7255169 89 + 28832112 GAUAUAUUCCUGGCAACGGGUAAUGCGAGUCUGCUGUCUUUUGUUGUGUGUG--CUGCUCGUUGAAUGCUUGUUGCCUGGACGAUGUUUGU (((((..(((.(((((((((((..((((((..((...(.........)...)--).))))))....))))))))))).)))..)))))... ( -31.20, z-score = -4.28, R) >droAna3.scaffold_13340 4043079 74 + 23697760 GAUAUAUUCCUGGCAACGAAUAAUGCGAGUCU-CUGUUGCCU-UUGUGUGU---------------UGCCGGUUGCCGUGGCAAUGUUUGU .........(((((((((.((((.((((....-...))))..-)))).)))---------------))))))((((....))))....... ( -21.60, z-score = -1.93, R) >consensus GAUAUAUUCCUGGCAACGAGUAAUGCGAGUCUGCUGCCUUUUGUUGUGUGUG__CUGCUCGUUGGAUGCUUGUUGCCUGGACGAUGUUUGU (((((..(((.(((((((((((..((((((..........................))))))....))))))))))).)))..)))))... (-21.33 = -22.92 + 1.59)
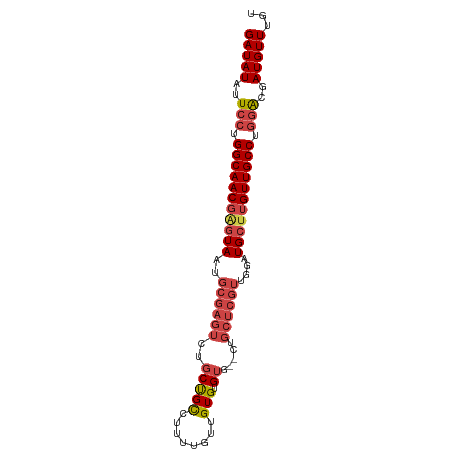

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:53:21 2011