| Sequence ID | dm3.chr3R |
|---|---|
| Location | 24,278,860 – 24,278,953 |
| Length | 93 |
| Max. P | 0.932517 |

| Location | 24,278,860 – 24,278,953 |
|---|---|
| Length | 93 |
| Sequences | 8 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 72.59 |
| Shannon entropy | 0.51787 |
| G+C content | 0.40091 |
| Mean single sequence MFE | -22.36 |
| Consensus MFE | -8.71 |
| Energy contribution | -9.14 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.39 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.37 |
| SVM RNA-class probability | 0.932517 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
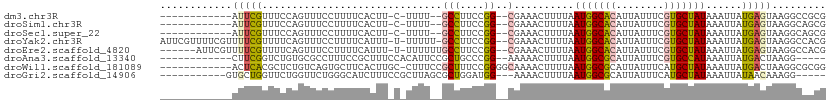
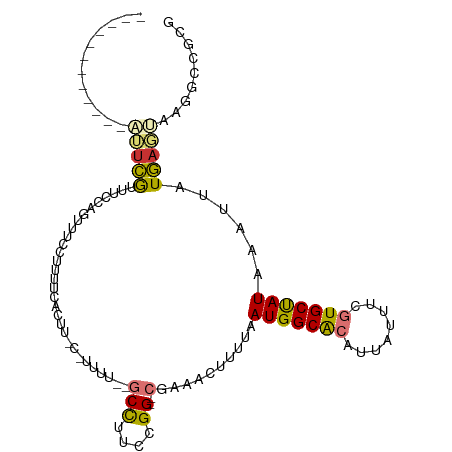
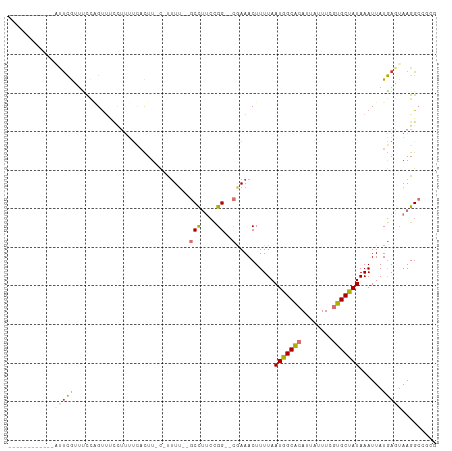
>dm3.chr3R 24278860 93 + 27905053 ------------AUUCGUUUCCAGUUUCCUUUUCACUU-C-UUUU--GCCUUCCGG--CGAAACUUUUAAUGGCACAUUAUUUCGUGCUAUAAAUUAUGAGUAAGGCCGCG ------------...........((..((((((((...-.-((((--(((....))--)))))......(((((((........)))))))......)))).))))..)). ( -22.50, z-score = -3.05, R) >droSim1.chr3R 23966741 93 + 27517382 ------------AUUCGUUUCCAGUUUCCUUUUCACUU-C-UUUU--GCCUUCCGG--CGAAACUUUUAAUGGCACAUUAUUUCGUGCUAUAAAUUAUGAGUAAGGCAGCG ------------...........(((.((((((((...-.-((((--(((....))--)))))......(((((((........)))))))......)))).)))).))). ( -23.90, z-score = -3.64, R) >droSec1.super_22 882353 93 + 1066717 ------------AUUCGUUUCCAGUUUCCUUUUCACUU-C-UUUU--GCCUUCCGG--CGAAACUUUUAAUGGCACAUUAUUUCGUGCUAUAAAUUAUGAGUAAGGCAGCG ------------...........(((.((((((((...-.-((((--(((....))--)))))......(((((((........)))))))......)))).)))).))). ( -23.90, z-score = -3.64, R) >droYak2.chr3R 6612453 106 - 28832112 AUUCGUUUUCGUUUUCGUUUUCAGUUUCCUUUUCAUUU-U-UUUUU-GCCUUCCGG--CGAAACUUUUAAUGGCACAUUAUUUCGUGCUAUAAAUUAUGAGUAAGGCCACG .......................((..(((((((((..-(-(((((-(((....))--)))).......(((((((........))))))))))..))))).))))..)). ( -23.30, z-score = -3.44, R) >droEre2.scaffold_4820 6709668 101 - 10470090 ------AUUCGUUUUCGUUUUCAGUUUCCUUUUCAUUU-U-UUUUUUGCCUUCCGG--CGAAACUUUUAAUGGCACAUUAUUUCGUGCUAUAAAUUAUGAGUAAGGCCACG ------.................((..(((((((((..-(-(((((((((....))--)))))......(((((((........))))))))))..))))).))))..)). ( -23.90, z-score = -4.03, R) >droAna3.scaffold_13340 17078586 92 + 23697760 ------------CUUCGGUCUGUGCGCCUUUCCGCUUUCCACAUUCCGCUGCCCGG--AAAAACUUUUAAUGGCGCAUUAUUUCGUGCCAUAAAUUAUGACUAAGG----- ------------(((..(((...(((......)))........(((((.....)))--)).........(((((((........))))))).......)))..)))----- ( -21.80, z-score = -1.41, R) >droWil1.scaffold_181089 8949518 98 + 12369635 ------------ACUCACGCUCUGUCAGUGCUUCACUUGC-CUUUCCGCUUUCCGGGGCAAAACUUUUAAUGGCGCAUUAUUUCAUGCUAUAAAUUAUGACUAAGGCGCGG ------------...(.((((..(((((((((....((((-(...(((.....))))))))..........))))).((((........))))....))))...)))).). ( -21.24, z-score = -0.30, R) >droGri2.scaffold_14906 1316306 92 + 14172833 -----------GUGCUGGUUCUGGUUCUGGGCAUCUUUCCGCUUAGCGCUGGAUGG---AAAACUUUUAAUGGCGCAUUAUUUCAUGCUAUAAAUUAUAACAAAGG----- -----------(((((...((..((.((((((........)))))).))..))(((---(.....))))..)))))..............................----- ( -18.30, z-score = 0.36, R) >consensus ____________AUUCGUUUCCAGUUUCCUUUUCACUU_C_UUUU__GCCUUCCGG__CGAAACUUUUAAUGGCACAUUAUUUCGUGCUAUAAAUUAUGAGUAAGGCCGCG ...............................................(((((..(........).....(((((((........)))))))...........))))).... ( -8.71 = -9.14 + 0.42)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:51:12 2011