
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 24,254,592 – 24,254,692 |
| Length | 100 |
| Max. P | 0.863729 |

| Location | 24,254,592 – 24,254,692 |
|---|---|
| Length | 100 |
| Sequences | 10 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 75.51 |
| Shannon entropy | 0.47186 |
| G+C content | 0.43265 |
| Mean single sequence MFE | -27.75 |
| Consensus MFE | -11.75 |
| Energy contribution | -10.98 |
| Covariance contribution | -0.77 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.25 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.863729 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 24254592 100 - 27905053 GAAUUUCCUUGCACGCAUUACGAAUCAAAUUU-ACGAAAAUUUAAUUUCUG--GCUCGCUUUCACGGGUCGCCAGAUCCUUGAAACGUGUGUGUGUGCGUGCU---------- ..........((((((((.(((((..((((((-....))))))...)))..--((.((((((((.(((((....))))).))))).))).)))))))))))).---------- ( -32.10, z-score = -3.24, R) >droSim1.chr3R 23942336 100 - 27517382 GAAUUUCCUUGCACGCAUUACGAAUCAAAUUU-ACGAAAAUUUAAUUUCUG--GCUCGCUUUCAUGGGUCGCCAGAUCCUUGAAACGUGUGUGUGUGCGUGCU---------- ..........((((((((.(((((..((((((-....))))))...)))..--((.((((((((.(((((....))))).))))).))).)))))))))))).---------- ( -32.10, z-score = -3.40, R) >droSec1.super_22 858272 100 - 1066717 GAAUUUCCUUGCACGCAUUACGAAUCAAAUUU-ACGAAAAUUUCAUUUCUG--GCUCGCUUUCAUGGGUCGCCAGAUCCUUGAAACGUGUGUGUGUGCGUGCU---------- ..........((((((((.(((((..((((((-....))))))...)))..--((.((((((((.(((((....))))).))))).))).)))))))))))).---------- ( -32.10, z-score = -3.19, R) >droEre2.scaffold_4820 6683004 106 + 10470090 GAAUUUCCUUGCACGCAUUACGAAUCAAAUUU-ACGAAAAUUUAAUUUCUG--GCUCGCUUUCAUGGGUCGCCAGAUCCUUGAAACGUGUGUGUAUGUGUGUGCGUGCU---- ..........((((((((((((((..((((((-....))))))...)))..--((.((((((((.(((((....))))).))))).))).))....))).)))))))).---- ( -32.60, z-score = -2.95, R) >droAna3.scaffold_13340 17054330 100 - 23697760 GAAUUUCCAAGCACGCAUUUCGAAUCAAAUUU-GCGAAAAUUUAAUUUCUG--GCUCACUUUCAUGGCUCGCCAAAUCCUGUGAAUGUGUGCGUGUGCGUGCC---------- ..........((((((((..((((..((((((-....))))))...)))..--((.(((.(((((((...........))))))).))).))).)))))))).---------- ( -27.00, z-score = -1.66, R) >dp4.chr2 20515513 96 + 30794189 GAAUUUCCACGCACGCAUUUCAAAUCAAAUUUGAGGAAAAUUUAAUUUCUGCCUCUCGCUUUCAUGGAUCGCCGGAU-------AUAUACACACACACACACU---------- .....(((......((...((((((...))))))(((((......))))).......))......((....))))).-------...................---------- ( -9.80, z-score = 0.54, R) >droWil1.scaffold_181089 8921323 91 - 12369635 GAAUUUCCACGCACGCAUUUCAAAUCAAAUUU-ACGAAAAUUUAAUUUCUG--GCUCGCUUUCAUGGCUCGCCAAAU---------GUGUGUGUGUGCGUGUU---------- .......(((((((((((..((..........-..((((......))))((--((..(((.....)))..))))..)---------)...)))))))))))..---------- ( -28.30, z-score = -3.52, R) >droVir3.scaffold_12822 3684211 103 - 4096053 GAAUUACCACGCACGCGUUUCAAAUCAAAUUU-ACGAAAAUUUAAUUUCAG--GCUCGCUUUCAUGCUCCGGCAAAU-------GUGCGUGCGUGUCCUCUGUGAUACGCAAC ........(((((((((..((((((.((((((-....)))))).))))..)--)..))).....(((....)))...-------))))))(((((((......)))))))... ( -27.60, z-score = -1.97, R) >droMoj3.scaffold_6540 16998061 103 - 34148556 GAAUUUCCACGCACGCGUUUCAAAUCAAAUUU-ACGAAAAUUUAAUUUCAG--GCUCGCUUUCAUGCUCCGGCGAAU-------GUGCGUGCGUGUCCCAUGUGAUACGCAAC ........(((((((.((((.((((.((((((-....)))))).)))).))--))(((((..........))))).)-------))))))(((((((......)))))))... ( -28.30, z-score = -1.58, R) >droGri2.scaffold_14906 1290595 103 - 14172833 GAAUUUCCACGCACGCGUUUCAAAUCAAAUUU-ACGAAAAUUUAAUUUCAG--GCUCGCUUUUAUGCUCCGGCAAAU-------GUGCGUGCGUGUCCCGUGUGAUACGCAAC ........(((((((((..((((((.((((((-....)))))).))))..)--)..))).....(((....)))...-------))))))(((((((......)))))))... ( -27.60, z-score = -1.48, R) >consensus GAAUUUCCACGCACGCAUUUCAAAUCAAAUUU_ACGAAAAUUUAAUUUCUG__GCUCGCUUUCAUGGGUCGCCAAAU_______AUGUGUGCGUGUGCGUGCU__________ ..........((((((.....((((.((((((.....)))))).))))................(((....)))............))))))..................... (-11.75 = -10.98 + -0.77)

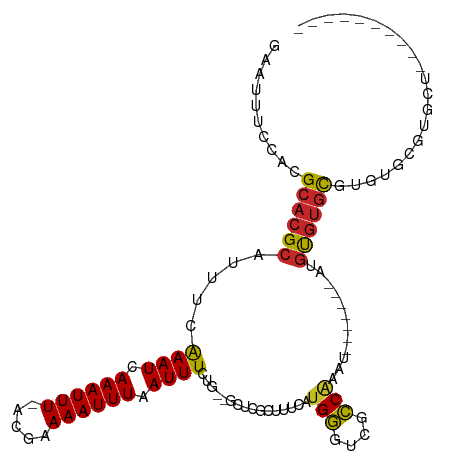

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:51:04 2011