
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 23,539,903 – 23,539,999 |
| Length | 96 |
| Max. P | 0.710221 |

| Location | 23,539,903 – 23,539,999 |
|---|---|
| Length | 96 |
| Sequences | 11 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 90.54 |
| Shannon entropy | 0.19647 |
| G+C content | 0.38492 |
| Mean single sequence MFE | -12.91 |
| Consensus MFE | -14.07 |
| Energy contribution | -14.17 |
| Covariance contribution | 0.10 |
| Combinations/Pair | 1.05 |
| Mean z-score | -0.16 |
| Structure conservation index | 1.09 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.47 |
| SVM RNA-class probability | 0.710221 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 23539903 96 + 27905053 ---UAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC-GGGCGCAAU-GACAAUAAGCACAAA-GCGACAAAUGAAGAGCCAAC ---.......((((.((((.(((((((....)))))))..............((.-..)))))))-))).....((.....-)).................. ( -11.40, z-score = 0.13, R) >droSim1.chr3R 23268885 96 + 27517382 ---UAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC-GGGCGCAAU-GACAAUAAGCACAAA-GCGACAAAUGAAGAGCCAAC ---.......((((.((((.(((((((....)))))))..............((.-..)))))))-))).....((.....-)).................. ( -11.40, z-score = 0.13, R) >droSec1.super_22 182595 96 + 1066717 ---UAAAAAGGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC-GGGCGCAAU-GACAAUAAGCACAAA-GCGACAAAUGAAGAGCCAAC ---......((((..(((..(((((((....)))))))..))).....((..((.-..(.((...-........)).)...-))..)).......))))... ( -13.80, z-score = -0.56, R) >droYak2.chr3R 5885178 96 - 28832112 ---UAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC-GGGCGCAAU-GACAAUAAGCACAAA-GCGACAAAUGAAGAGCCAAC ---.......((((.((((.(((((((....)))))))..............((.-..)))))))-))).....((.....-)).................. ( -11.40, z-score = 0.13, R) >droEre2.scaffold_4820 5974257 84 - 10470090 ---UAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC-GGGCGCAAU-GACAAUAAGCACAAA-GCGACAAA------------ ---.......((((.((((.(((((((....)))))))..............((.-..)))))))-))).....((.....-))......------------ ( -11.40, z-score = -0.52, R) >droAna3.scaffold_13340 4549099 96 - 23697760 ---UAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAGGAAAAUGAAGCC-GGGCGCAGU-GACAAUAAGCACAAA-GCGACAAAUGAAGAGCCAAC ---.......((((.((((.(((((((....)))))))....((.........))-....)))))-))).....((.....-)).................. ( -11.90, z-score = 0.52, R) >dp4.chr2 3900076 93 + 30794189 ---UAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC-GGGCGCAAU-GACAAUCAGCACAAAAGCGACAAAUGGAGAGA---- ---.......((((.((((.(((((((....)))))))..............((.-..)))))))-))).....((......))..............---- ( -11.60, z-score = 0.22, R) >droPer1.super_7 3994552 93 - 4445127 ---UAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC-GGGCGCAAU-GACAAUCAGCACAAAAGCGACAAAUGGAGAGA---- ---.......((((.((((.(((((((....)))))))..............((.-..)))))))-))).....((......))..............---- ( -11.60, z-score = 0.22, R) >droMoj3.scaffold_6540 1788006 99 - 34148556 CUGAAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC-CAGCGCCGUUGACAAUAAGCGCCAA-GCG-CAAAUGAAGAGCCAGC (((.......((((((.((.(((((((....)))))))..............((.-..)))).)))))).....((((...-)))-)...........))). ( -17.90, z-score = -0.93, R) >droGri2.scaffold_14830 5832736 98 - 6267026 ---AAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCCAGGGCGCACUUGACAAUAAGCGCAAA-GCGACAAAUGAAGAGCCACC ---.......(((((.(((.(((((((....)))))))..............((....))))).))))).....((.....-)).................. ( -16.10, z-score = -0.73, R) >droVir3.scaffold_12855 8098893 96 - 10161210 ---AAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC-CAGCGCCGUUGACAAUAAGCGCCAA-GCGACAAAUGAAGAGCCAA- ---.......(((..(((..(((((((....)))))))..))).........((.-..((((............))))...-)))))..............- ( -13.50, z-score = -0.35, R) >consensus ___UAAAAAAGUUAAUUGCAGUUAAUGACGACAUUAAUCACAAGAAAAUGAAGCC_GGGCGCAAU_GACAAUAAGCACAAA_GCGACAAAUGAAGAGCCAAC ..........(((.(((((.(((((((....)))))))..............((....))))))).))).....((......)).................. (-14.07 = -14.17 + 0.10)

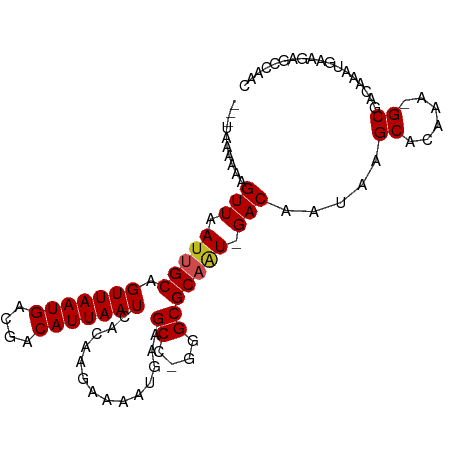
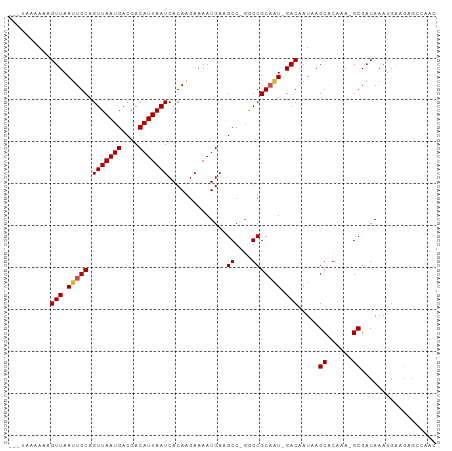
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:48:57 2011