
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 23,524,507 – 23,524,627 |
| Length | 120 |
| Max. P | 0.670228 |

| Location | 23,524,507 – 23,524,627 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 75.83 |
| Shannon entropy | 0.34499 |
| G+C content | 0.31786 |
| Mean single sequence MFE | -26.71 |
| Consensus MFE | -14.50 |
| Energy contribution | -16.51 |
| Covariance contribution | 2.01 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.670228 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 23524507 120 + 27905053 UCCAUUACUCUAGAGCCAUUGAUUGGUACAUUAUUUUUGUUGAAUGGAAUGAUUCAUUUCAAUCGUUCUUAUCAUAUAAUUUCAAAGGAAAUAACUGGGUAAUGUCCCCACGUAUCAAAA ..((((((.((((.((((.....))))...(((((((..(((((.(((((((((......)))))))))...........)))))..)))))))))))))))))................ ( -28.20, z-score = -2.31, R) >droEre2.scaffold_4820 5958934 112 - 10470090 UCCAUUUCUUUUGAGCAAGUGAUUGGAAC--------UGUUGAAUUGAAUGAUUCAUUCCGAUUGUUCUUAUCAUAUGAUUUAAAAGAAAAUAAUAGGGUAAUGAACCCACGUAACAAAA ....(((((((((((((.(((((.(((((--------.((((...(((.....)))...)))).))))).))))).)).)))))))))))......((((.....))))........... ( -30.30, z-score = -3.74, R) >droYak2.chr3R 5869583 114 - 28832112 UCGAUUUUUUUUAAGACAGUGAUCGGAACUUUAUCUCUGUUGAAUUGAAUGAUUCAUUC------UUCUUAUCAUAUAAUUUGAACGGAAAUAAUCGGGUCAUGUACCCGCGAAACCAAA ..((((..............))))((....((((.((((((((((((.(((((......------.....))))).)))))).)))))).)))).(((((.....))))).....))... ( -21.64, z-score = -0.17, R) >consensus UCCAUUUCUUUUGAGCCAGUGAUUGGAAC_UUAU_U_UGUUGAAUUGAAUGAUUCAUUCC_AU_GUUCUUAUCAUAUAAUUUAAAAGGAAAUAAUAGGGUAAUGUACCCACGUAACAAAA ....(((((((((((...(((((.(((((...........((((((....))))))........))))).)))))....)))))))))))......(((.......)))........... (-14.50 = -16.51 + 2.01)

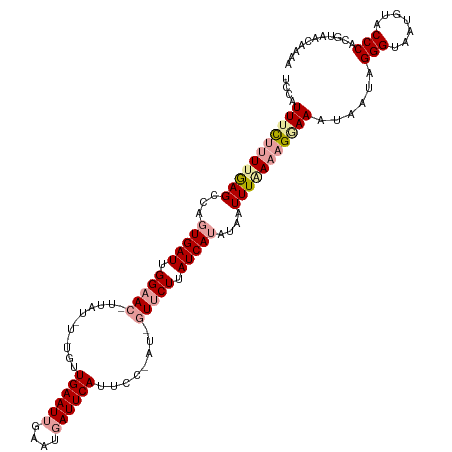

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:48:56 2011