
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 23,334,423 – 23,334,584 |
| Length | 161 |
| Max. P | 0.580801 |

| Location | 23,334,423 – 23,334,584 |
|---|---|
| Length | 161 |
| Sequences | 6 |
| Columns | 180 |
| Reading direction | forward |
| Mean pairwise identity | 69.53 |
| Shannon entropy | 0.55323 |
| G+C content | 0.38547 |
| Mean single sequence MFE | -40.27 |
| Consensus MFE | -15.28 |
| Energy contribution | -14.68 |
| Covariance contribution | -0.60 |
| Combinations/Pair | 1.61 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.580801 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 23334423 161 + 27905053 AAUGCGCAGUUCGCAUAGGGAACU---------UAACC-----UUAUUACCGGAACUAACAGGUGCUUGUACUUUCCUAUUUUGGAAUGCAUAAACAGUGC---UUUAAUUAAA-CACUUUUUCACUCUCAAGUGAAGGAAGUGCAUUUUU-CUAAGUGCACGAAACCCGUGCUCUUCAG ((((((((((((..((((((....---------...))-----)))).....)))))...(((..((((((..((((......)))))))).....))..)---))........-...(((((((((....))))))))).)))))))...-..(((.(((((.....))))).)))... ( -41.00, z-score = -2.00, R) >droSim1.chr3R 23071903 159 + 27517382 AAAGAGCAGUUCGCAUAGGGAACU---------UAACC-----UGAUUACCGGAACUAACAGGUGCUUGCACUUUCCUAUCUUGAAAUGUAUAAAUAGUGC---UUUAAUUAAA-CACUUUUUC--UCGCAAGUGAAGGAAGUGCAUUUUU-CUAAGUGCACGAAACCCGUGCUCUUCAG .(((((((((((...((((.....---------...))-----)).......))))).((.(((...(((((((.........(((((((((.....(((.---(((....)))-)))((((((--.(....).)))))).))))))))).-..)))))))....))).))))))))... ( -39.80, z-score = -1.39, R) >droSec1.super_13 2077689 159 + 2104621 AAUGAGCAGUUCGCAUAGGGAACU---------UAACC-----UGAUUACCAGAACUAACAGGUGCUUGCACUUUCCUAUCUUGAACUGUAUAAAUAGUGC---CUUAAUUAAA-CACUUUUUC--UCUCAAGUGAAGGAAGUGCAUUUUU-CUAAGUGCACGAAACCCGUGCUCUUCAG ...(((((((((.(...).)))))---------..(((-----((.(((.......))))))))))))((((((.(((.......(((((....)))))..---..........-(((((....--....))))).)))))))))......-..(((.(((((.....))))).)))... ( -39.80, z-score = -1.78, R) >droYak2.chr3R 5683684 175 - 28832112 AAUGGGCAGUUCACAUAGAGAGCUGUCUAAGCUUAACUCACCUUGAUUACCGAAACUAACAGGUGUUUGUACUUGCGGAACUUGAAAUGCUCGUGUAGUGC---UUUAAUUAAAGCACUUUUCC--UCUCGAGUGUAGGAAGUGCAUUUUUUCUAAGUGCACAAAACCCCUGCUCUUCAG ..((((((((((.......)))))))))).((...((.(((((.(.(((.......)))))))))...))....))......((((..(((((.(.(((((---(((....)))))))).....--.).)))))(((((..((((((((.....))))))))......)))))..)))). ( -49.50, z-score = -1.77, R) >droEre2.scaffold_4820 5773617 165 - 10470090 AAUGCCCAGUUCGCAUAGAGAACU---------UAACCCAACUUUAUUAGCGAAACUAGCAGUUGUUUGUGCCUGCGGAUCUUGAAAUGCAUGUGUAGUGC---CCUAAUUAAAACACUUUUUC--UCUCAAGUGAGGGAAGUGUAUUUUU-CUAAGUGCACGAAACCCCUGCUCUUCAG .......(((((((((((((....---------........))))))..))).))))(((((...(((((((((..(((....((((((((((((((((..---....)))...))))...(((--((((....)))))))))))))))))-)).)).)))))))....)))))...... ( -42.20, z-score = -1.74, R) >droAna3.scaffold_13340 4355429 160 - 23697760 -------AGUAUUUUGUGAGCAUU-------UAUGAAU---UUUGAGUCCACUAACCCAAAAGUUUUUGAAGUUUUUUAAAGCAAAAUUUGUGUAUGAUUCAAUUUUGAUUUAA--UCUUUUGUAAAUAAAAUAAAAUCGCACUUAUUUUUCGCGAGUGUAGGAACCUCCAACCACUCU- -------........((((((.((-------((.((((---((..((...(((........))).))..))))))..))))))(((((..((((..((((..((((((.((((.--.......))))))))))..))))))))..)))))))))(((((..((.....))...))))).- ( -29.30, z-score = -1.37, R) >consensus AAUGAGCAGUUCGCAUAGAGAACU_________UAACC_____UGAUUACCGGAACUAACAGGUGCUUGUACUUUCCUAUCUUGAAAUGCAUAAAUAGUGC___UUUAAUUAAA_CACUUUUUC__UCUCAAGUGAAGGAAGUGCAUUUUU_CUAAGUGCACGAAACCCCUGCUCUUCAG ............((((.(((...................................................(((((.....(((((..((((.....))))...))))).......((((..........)))))))))..((((((((.....))))))))....)))))))....... (-15.28 = -14.68 + -0.60)

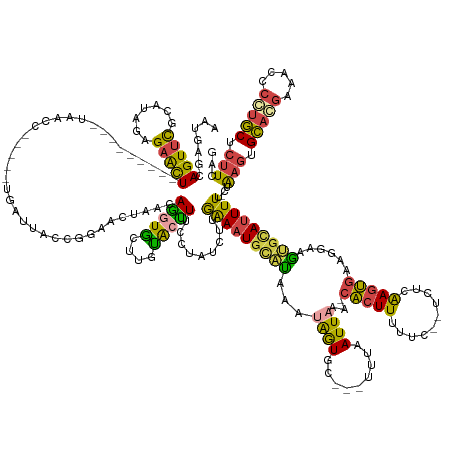

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:48:19 2011