
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 23,107,971 – 23,108,074 |
| Length | 103 |
| Max. P | 0.821891 |

| Location | 23,107,971 – 23,108,074 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 76.63 |
| Shannon entropy | 0.43579 |
| G+C content | 0.35866 |
| Mean single sequence MFE | -22.07 |
| Consensus MFE | -11.10 |
| Energy contribution | -11.72 |
| Covariance contribution | 0.62 |
| Combinations/Pair | 1.39 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.821891 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
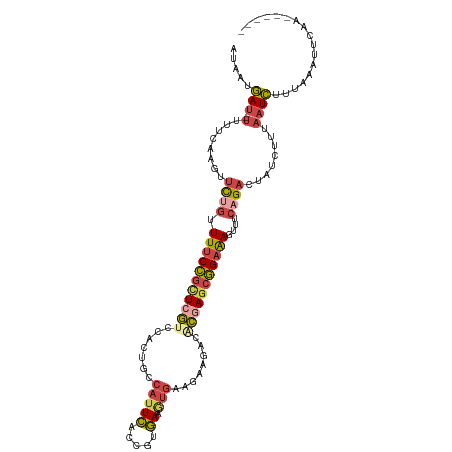
>dm3.chr3R 23107971 103 + 27905053 AUAAUGAU--UUUUCAAGUUUUGUUUUCCGCUCGUC---UGCCAUUCUCCGUGAAGUGAAGAAGACUCUAACGGAAAGUUUCAGACUAUCUAUAAUCUUUAAAUUCAA------ ....(((.--...))).................(((---((..(((.(((((..(((.......)))...))))).)))..)))))......................------ ( -16.10, z-score = 0.19, R) >droAna3.scaffold_13340 4114613 99 - 23697760 AUAAUAAUGAUUUUCAAGUUUUUUUUUCAUUUCAUUCAUUGCCAUUCGUCAUGA---------UUCGUAAGCUGCGACUUUCCGUAUUCAUUUUUUUACGAGAAUCGU------ ......(((((((((.(((((.....((((..(..............)..))))---------.....)))))(((......)))..............)))))))))------ ( -13.34, z-score = -0.52, R) >droEre2.scaffold_4820 5544013 106 - 10470090 AUAAUGAU--UUUUCAAGUUCUGUUUUCCGCUCGUCCACUGCCAUUCACCGUGAAAUGAAGAAGACACGAGCGGAAAGUUUCAGACUAUCUCUAAUCUACAGAUCCAA------ .....(((--(.....(((.(((((((((((((((...((....((((........))))..))..))))))))))))...))))))......))))...........------ ( -25.00, z-score = -1.85, R) >droYak2.chr3R 5446462 112 - 28832112 AUAAUGAU--UUUUCAAGUUCUGUUUUCCGCUCGUCCACUGCCAUUUACCGUAAAAUGAAAAAUACACGAGCGGAAACUUUCAGACUAUCUCUAAUCUUUAUCUUUAUAUCCAA ((((.(((--(.....(((.(((.(((((((((((.......(((((......)))))........)))))))))))....))))))......)))).))))............ ( -24.56, z-score = -4.58, R) >droSec1.super_13 1862048 106 + 2104621 AUAAUGAU--UUUUCAAGUUCUGUUUUCCGCUCGUCCACUGCCAUUCACCGUGAAGUGAAGAAGACACGAGCGGAAAGUUUCAGACUAUCUUUAAUCUUCAAAUUCAA------ .....(((--(.....(((.(((((((((((((((...((....(((((......)))))..))..))))))))))))...))))))......))))...........------ ( -28.30, z-score = -3.05, R) >droSim1.chr3R_random 36634 106 + 1307089 AUAAUGAU--UUUUCAAGUUCUGUUUUCCGCUCGUCCACUGCCAUUCACCGUGAAGUGAAGAAGACUCGAGCGGAAAGUUUCUGACUAUCUUUAAUCUUUAAAUUCAA------ .(((.(((--(.....((((...(((((((((((....((....(((((......)))))..))...))))))))))).....))))......)))).))).......------ ( -25.10, z-score = -2.06, R) >consensus AUAAUGAU__UUUUCAAGUUCUGUUUUCCGCUCGUCCACUGCCAUUCACCGUGAAGUGAAGAAGACACGAGCGGAAAGUUUCAGACUAUCUUUAAUCUUUAAAUUCAA______ ...................((((.(((((((((((.........(((.....)))...........)))))))))))....))))............................. (-11.10 = -11.72 + 0.62)



| Location | 23,107,971 – 23,108,074 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 76.63 |
| Shannon entropy | 0.43579 |
| G+C content | 0.35866 |
| Mean single sequence MFE | -24.53 |
| Consensus MFE | -13.07 |
| Energy contribution | -13.88 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.619974 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 23107971 103 - 27905053 ------UUGAAUUUAAAGAUUAUAGAUAGUCUGAAACUUUCCGUUAGAGUCUUCUUCACUUCACGGAGAAUGGCA---GACGAGCGGAAAACAAAACUUGAAAA--AUCAUUAU ------.(((.(((..((((((....)))))).....((((((((...((((.((...(((....)))...)).)---))).))))))))...........)))--.))).... ( -21.20, z-score = -1.13, R) >droAna3.scaffold_13340 4114613 99 + 23697760 ------ACGAUUCUCGUAAAAAAAUGAAUACGGAAAGUCGCAGCUUACGAA---------UCAUGACGAAUGGCAAUGAAUGAAAUGAAAAAAAAACUUGAAAAUCAUUAUUAU ------.(((((.(((((..........)))))..)))))..(((..((..---------......))...))).........(((((................)))))..... ( -8.39, z-score = 1.56, R) >droEre2.scaffold_4820 5544013 106 + 10470090 ------UUGGAUCUGUAGAUUAGAGAUAGUCUGAAACUUUCCGCUCGUGUCUUCUUCAUUUCACGGUGAAUGGCAGUGGACGAGCGGAAAACAGAACUUGAAAA--AUCAUUAU ------..((.(((((((((((....)))))).....(((((((((((.((((((((((......))))).)).)).).)))))))))))))))).))......--........ ( -31.50, z-score = -2.26, R) >droYak2.chr3R 5446462 112 + 28832112 UUGGAUAUAAAGAUAAAGAUUAGAGAUAGUCUGAAAGUUUCCGCUCGUGUAUUUUUCAUUUUACGGUAAAUGGCAGUGGACGAGCGGAAAACAGAACUUGAAAA--AUCAUUAU ............((((.((((.(((....((((....(((((((((((.((((..((((((......)))))).)))).))))))))))).)))).)))....)--))).)))) ( -31.10, z-score = -3.90, R) >droSec1.super_13 1862048 106 - 2104621 ------UUGAAUUUGAAGAUUAAAGAUAGUCUGAAACUUUCCGCUCGUGUCUUCUUCACUUCACGGUGAAUGGCAGUGGACGAGCGGAAAACAGAACUUGAAAA--AUCAUUAU ------...........((((......((((((....(((((((((((.(((((((((((....)))))).)).)).).))))))))))).))).))).....)--)))..... ( -29.70, z-score = -2.40, R) >droSim1.chr3R_random 36634 106 - 1307089 ------UUGAAUUUAAAGAUUAAAGAUAGUCAGAAACUUUCCGCUCGAGUCUUCUUCACUUCACGGUGAAUGGCAGUGGACGAGCGGAAAACAGAACUUGAAAA--AUCAUUAU ------.(((.(((...(((((....)))))......((((((((((...((((((((((....)))))).)).))....))))))))))...........)))--.))).... ( -25.30, z-score = -1.40, R) >consensus ______UUGAAUUUAAAGAUUAAAGAUAGUCUGAAACUUUCCGCUCGUGUCUUCUUCACUUCACGGUGAAUGGCAGUGGACGAGCGGAAAACAGAACUUGAAAA__AUCAUUAU ................((((((....)))))).....((((((((((............(((((.((.....)).)))))))))))))))........................ (-13.07 = -13.88 + 0.81)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:47:38 2011