
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 8,762,113 – 8,762,203 |
| Length | 90 |
| Max. P | 0.538535 |

| Location | 8,762,113 – 8,762,203 |
|---|---|
| Length | 90 |
| Sequences | 8 |
| Columns | 104 |
| Reading direction | reverse |
| Mean pairwise identity | 79.34 |
| Shannon entropy | 0.37822 |
| G+C content | 0.49054 |
| Mean single sequence MFE | -22.64 |
| Consensus MFE | -12.59 |
| Energy contribution | -13.12 |
| Covariance contribution | 0.53 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.538535 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
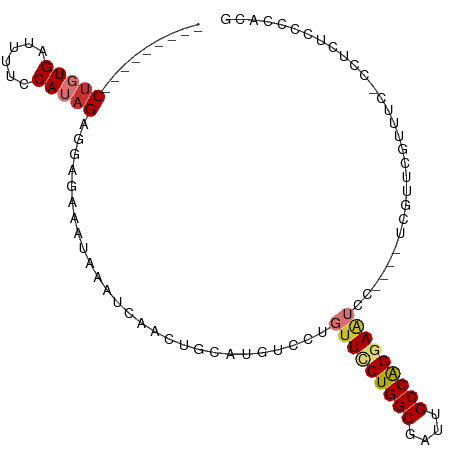
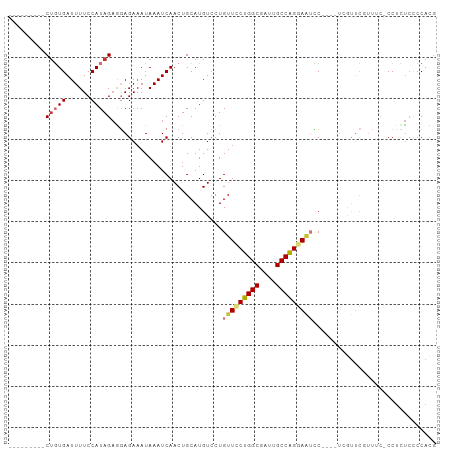
>dm3.chr2L 8762113 90 - 23011544 ---------CUGUGAUUUUCCAUAGAGGAGAAAUAAAUCAACUGCAUGUCCUGUUUCUGGCGAUUGCCAGGAAUUC----UGGUUCGUUUC-CCUCUCCCCACG ---------.((((......))))((((.((((..(((((((.....))...(((((((((....)))))))))..----)))))..))))-))))........ ( -25.50, z-score = -2.33, R) >droSim1.chr2L 8540857 90 - 22036055 ---------CUGUGAUUUUCCAUAGAGGAGAAAUAAAUCAACUGCAUGUCCUGUUUCUGGCGAUUGCCAGGAAUCC----UGGUUCGUUUC-CCUCUCCCCACG ---------.((((......))))((((.((((..(((((((.....))...(((((((((....)))))))))..----)))))..))))-))))........ ( -25.50, z-score = -2.33, R) >droSec1.super_3 4235497 90 - 7220098 ---------CUGUGAUUUUCCAUAGAGGAGAUAUAAAUCAACUGCAUGUCCUGUUUCUGGCGAUUGCCAGGAAUCC----UGGUUCGUUUC-CCUCUCCCCACG ---------(((((......))))).(((((...((((.(((((((.....)))(((((((....)))))))....----.)))).)))).-..)))))..... ( -22.90, z-score = -1.40, R) >droYak2.chr2L 11405061 94 - 22324452 ---------CUGUGAUUUUCCAUAGAGGAGAAAUAAAUCAACUGCAUGUCCUGUUCCUGGCGAUUGCCAGGAAUCCUCGUUCGUUCGUUCC-CCUCUCCCCACG ---------(((((......))))).(((((....(((.(((.(.(((....(((((((((....)))))))))...))).)))).)))..-..)))))..... ( -27.00, z-score = -3.23, R) >droEre2.scaffold_4929 9349049 90 - 26641161 ---------CUGUGAUUUUCCAUAGCGGAGAAAUAAAUCAACUGCAUGUCCUGUUUCUGGCGAUUGCCAGGAAUCC----UCCUUCGUUGC-CCCUCCUCCGUG ---------.((((......))))((((((........((((..........(((((((((....)))))))))..----......)))).-.....)))))). ( -22.21, z-score = -1.85, R) >droAna3.scaffold_12916 10401845 98 - 16180835 CUCAACGCUCCGUGAUUUUCCAUAGAAGAGAAAUAAAUCAACUGCAUGUCCUGUUCCUGGCGAUUGCCAGGACUCC----UCAUUCUUUUCUCCUCUGCCCG-- ......((...(((......)))(((.((((((..(((.....(((.....)))(((((((....)))))))....----..)))..)))))).)))))...-- ( -21.20, z-score = -1.63, R) >dp4.chr4_group5 1547722 89 + 2436548 --CUCCGCUCUUUGAUUUUCCAUAGAAUAGAAAUAAAUCAACUGCAUGUCCUGUUCCUGGCGAUUGCCGGUAGGGC-----CAC--------CCCGUCCCCGUG --....((...(((((((................)))))))..))..((((((...(((((....)))))))))))-----...--------............ ( -18.39, z-score = -0.22, R) >droPer1.super_5 1516537 89 + 6813705 --CUCCGCUCUUUGAUUUUCCAUAGAAUAGAAAUAAAUCAACUGCAUGUCCUGUUCCUGGCGAUUGCCGGUAGGGC-----CAC--------CCCUUCCCCGUG --....((...(((((((................)))))))..))..((((((...(((((....)))))))))))-----...--------............ ( -18.39, z-score = -0.49, R) >consensus _________CUGUGAUUUUCCAUAGAGGAGAAAUAAAUCAACUGCAUGUCCUGUUCCUGGCGAUUGCCAGGAAUCC____UCGUUCGUUUC_CCUCUCCCCACG .........(((((......)))))...........................(((((((((....))))))))).............................. (-12.59 = -13.12 + 0.53)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:26:54 2011