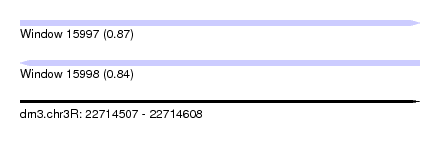

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 22,714,507 – 22,714,608 |
| Length | 101 |
| Max. P | 0.867561 |

| Location | 22,714,507 – 22,714,608 |
|---|---|
| Length | 101 |
| Sequences | 7 |
| Columns | 107 |
| Reading direction | forward |
| Mean pairwise identity | 69.76 |
| Shannon entropy | 0.61407 |
| G+C content | 0.47787 |
| Mean single sequence MFE | -30.33 |
| Consensus MFE | -12.15 |
| Energy contribution | -14.21 |
| Covariance contribution | 2.06 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.867561 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 22714507 101 + 27905053 UGGAAACUUGACGAGAAUACAGCUGCAGAUGCGGGCAGAUGCAGAUGCAGAUGCACAU----AUAUGUAUGUAGAUGCAGAGCAUGUGCAGCCUCAACGA--AAUAU .(....)(((..(((......((((((.((((..(((..(((....)))..)))....----...(((((....)))))..)))).)))))))))..)))--..... ( -32.00, z-score = -2.44, R) >droSim1.chr3R 22545405 86 + 27517382 UGAAAACUUGACGAGAAUACAGCUGCAGAUGCGGGCAGAUGCAGAUGCA------------------UCUGUAGAUGCAGAGCAUGUGCAGCCGCAACGA--AUGA- .....................((((((.((((..(((..((((((....------------------))))))..)))...)))).))))))........--....- ( -31.00, z-score = -3.00, R) >droSec1.super_13 1487787 102 + 2104621 UGAAAACUUGACGAGAAUACAGCUGCAGAUGCGGGCAGAUGCAGAUGCAGAUACAGAUGC--ACAUACAUGUGGAUGCAGAGCAUGUGCAGCCGCAACGA--AUGA- .....................((((((.((((..(((..((((..((((........)))--)......))))..)))...)))).))))))........--....- ( -29.00, z-score = -1.32, R) >droYak2.chr3R 5057995 101 - 28832112 UGGAAACUUGACGAGUAUACAGCUGCAGAUGCGGGCAGAUGCAGAUGCAGAUACAUAGGC--ACAUAUAUGUAGAUGCAGAGCAUGUGCAGGCACAACAACGA---- .(....)....((.........((((....))))(((.((((...((((..((((((...--.....))))))..))))..)))).)))...........)).---- ( -29.70, z-score = -2.26, R) >droEre2.scaffold_4820 5144165 106 - 10470090 UGGAAACUUGACGAGAAUACAGCUGCAGAUGCGGGCAGAUGCAGAUGCAGAUACAGAUACAGAUGCACAUAUAGAUGCAGAGCAUGUGCAUGUGCAGCCACAACGA- (((...................(((((..((((......))))..))))).............(((((((.((.((((...)))).)).))))))).)))......- ( -29.40, z-score = -1.47, R) >droMoj3.scaffold_6540 11710523 96 + 34148556 -UGAAACUUG----GGUUGUGU-UGCAG-CACCCGC--AUGCAGAUGCAGAUGCAGAUGGUGAUGUCGAUGUAGAUAAAAUUCAUGUGCAGCGACUGC-AUAAUAA- -........(----(((.((..-....)-)))))..--((((((.(((...((((.((((...((((......))))....)))).))))))).))))-)).....- ( -28.90, z-score = -0.90, R) >droVir3.scaffold_12855 2499847 103 + 10161210 -UGAAACUUGUACGGGAUGCGGGCGCAG-CAGAUGCCGAUGCUGAUGCAGAUGCAGAUGUCGAAGUCGAUGUAGAUAAAGCUCAUGUGCAUCGGCUGC-AUAAUAA- -...............((((((((((((-((........))))).))).((((((.(((.(...(((......)))...)..))).))))))..))))-)).....- ( -32.30, z-score = -0.41, R) >consensus UGGAAACUUGACGAGAAUACAGCUGCAGAUGCGGGCAGAUGCAGAUGCAGAUACAGAUGC__AUAUAUAUGUAGAUGCAGAGCAUGUGCAGCCGCAACGA__AUGA_ ......................(((((.((((..((....))...((((..((((..............))))..))))..)))).)))))................ (-12.15 = -14.21 + 2.06)
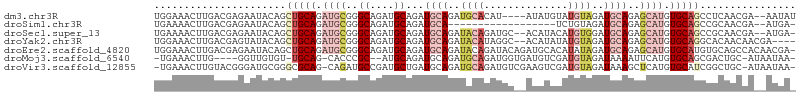


| Location | 22,714,507 – 22,714,608 |
|---|---|
| Length | 101 |
| Sequences | 7 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 69.76 |
| Shannon entropy | 0.61407 |
| G+C content | 0.47787 |
| Mean single sequence MFE | -27.46 |
| Consensus MFE | -10.82 |
| Energy contribution | -11.16 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.840887 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 22714507 101 - 27905053 AUAUU--UCGUUGAGGCUGCACAUGCUCUGCAUCUACAUACAUAU----AUGUGCAUCUGCAUCUGCAUCUGCCCGCAUCUGCAGCUGUAUUCUCGUCAAGUUUCCA .....--.......(((((((.((((.........(((((....)----))))(((..(((....)))..)))..)))).))))))).................... ( -24.90, z-score = -0.76, R) >droSim1.chr3R 22545405 86 - 27517382 -UCAU--UCGUUGCGGCUGCACAUGCUCUGCAUCUACAGA------------------UGCAUCUGCAUCUGCCCGCAUCUGCAGCUGUAUUCUCGUCAAGUUUUCA -....--.((.((((((((((.((((..((((((....))------------------))))...((....))..)))).))))))))))....))........... ( -30.20, z-score = -3.45, R) >droSec1.super_13 1487787 102 - 2104621 -UCAU--UCGUUGCGGCUGCACAUGCUCUGCAUCCACAUGUAUGU--GCAUCUGUAUCUGCAUCUGCAUCUGCCCGCAUCUGCAGCUGUAUUCUCGUCAAGUUUUCA -....--.((.((((((((((.((((...(((.....(((((.((--(((........))))).))))).)))..)))).))))))))))....))........... ( -32.00, z-score = -2.67, R) >droYak2.chr3R 5057995 101 + 28832112 ----UCGUUGUUGUGCCUGCACAUGCUCUGCAUCUACAUAUAUGU--GCCUAUGUAUCUGCAUCUGCAUCUGCCCGCAUCUGCAGCUGUAUACUCGUCAAGUUUCCA ----..(((((.((((..(((.((((..((((..((((((.....--...))))))..))))...)))).)))..))))..)))))..................... ( -28.00, z-score = -2.07, R) >droEre2.scaffold_4820 5144165 106 + 10470090 -UCGUUGUGGCUGCACAUGCACAUGCUCUGCAUCUAUAUGUGCAUCUGUAUCUGUAUCUGCAUCUGCAUCUGCCCGCAUCUGCAGCUGUAUUCUCGUCAAGUUUCCA -.((.((..((((((.((((..((((..((((..((((.((((....)))).))))..))))...))))......)))).))))))..))....))........... ( -30.40, z-score = -1.63, R) >droMoj3.scaffold_6540 11710523 96 - 34148556 -UUAUUAU-GCAGUCGCUGCACAUGAAUUUUAUCUACAUCGACAUCACCAUCUGCAUCUGCAUCUGCAU--GCGGGUG-CUGCA-ACACAACC----CAAGUUUCA- -......(-(((((((.((...((((...))))...)).))))..........((((((((((....))--)))))))-)))))-........----.........- ( -22.60, z-score = -0.99, R) >droVir3.scaffold_12855 2499847 103 - 10161210 -UUAUUAU-GCAGCCGAUGCACAUGAGCUUUAUCUACAUCGACUUCGACAUCUGCAUCUGCAUCAGCAUCGGCAUCUG-CUGCGCCCGCAUCCCGUACAAGUUUCA- -.......-(((((.(((((....(((((.........(((....)))....(((....)))..))).)).))))).)-)))).......................- ( -24.10, z-score = -0.27, R) >consensus _UCAU__UCGUUGCGGCUGCACAUGCUCUGCAUCUACAUAUACAU__GCAUCUGCAUCUGCAUCUGCAUCUGCCCGCAUCUGCAGCUGUAUUCUCGUCAAGUUUCCA .........(((((((.(((..((((...))))....................((...(((....)))...))..))).)))))))..................... (-10.82 = -11.16 + 0.33)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:46:49 2011