
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 22,644,629 – 22,644,723 |
| Length | 94 |
| Max. P | 0.739727 |

| Location | 22,644,629 – 22,644,723 |
|---|---|
| Length | 94 |
| Sequences | 11 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 71.74 |
| Shannon entropy | 0.56126 |
| G+C content | 0.38590 |
| Mean single sequence MFE | -14.98 |
| Consensus MFE | -8.86 |
| Energy contribution | -8.86 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.75 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.645220 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3R 22644629 94 + 27905053 -----------------UAUGUACAUAUAUGUGCUUUGCGCUUUCCUUACCUCUCCGCAAAUGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG -----------------...((((..........((((((...............)))))).(((((((((...............)))))))))........)))).... ( -16.72, z-score = -1.74, R) >droSim1.chr3R 22475785 94 + 27517382 -----------------UACAUAUAUGCAUGUGCUUUUCGCUUUCCUUACCUCUCCGCAAAUGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG -----------------.............(((((....((...............))....(((((((((...............))))))))).......))))).... ( -12.92, z-score = -0.54, R) >droEre2.scaffold_4820 5071283 94 - 10470090 -----------------CACAUAUGUAUACGUGCUUUUCGCUAUUCUUACCUCUCCGCAAAUGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG -----------------.............(((((....(((((((.......(((((....))))).....)))))....))((......)).........))))).... ( -15.00, z-score = -1.35, R) >droYak2.chr3R 4984041 94 - 28832112 -----------------UAUGUACAUAUAUGUACUUUUCGCUUUCCAUACCUCUCCGCAAAUGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG -----------------...(((((....)))))...((((............(((((....)))))..............))))........((((....))))...... ( -14.17, z-score = -1.32, R) >droSec1.super_13 1418411 94 + 2104621 -----------------UACAUAUAUGUAUGUGCUUUUCGCUUUCCUUACCUCUCCGCAAAUGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG -----------------((((....)))).(((((....((...............))....(((((((((...............))))))))).......))))).... ( -13.32, z-score = -0.92, R) >droWil1.scaffold_181130 13966757 73 - 16660200 --------------------------------------GUGUGGGUGUGUGUGUUUGUGUGUGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG --------------------------------------(((.((((((....((((.(......).))))....)))))))))((......))((((....))))...... ( -10.80, z-score = 0.07, R) >dp4.chr2 25963407 86 - 30794189 -----------------------CACACACAUGCUCUGGCCUCACCUU-CGACGCCGCAA-UGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG -----------------------........(((...(((.((.....-.)).)))))).-.(((((((((...............)))))))))................ ( -15.86, z-score = -0.94, R) >droPer1.super_19 1340670 86 - 1869541 -----------------------CACACACAUGCUCUGGCCUCACCUU-CGACGCCGCAA-UGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG -----------------------........(((...(((.((.....-.)).)))))).-.(((((((((...............)))))))))................ ( -15.86, z-score = -0.94, R) >droAna3.scaffold_13340 7008992 85 - 23697760 -------------------------UUUAUGUACGUAUGUAUGUCCUUACCUCUCCGCAA-UGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG -------------------------.....((((....(((......)))..........-.(((((((((...............)))))))))........)))).... ( -13.06, z-score = -0.97, R) >droVir3.scaffold_13047 11963302 102 + 19223366 --------UACAUAUGCCACAGCAGGAGUUGCCACAACCACUCACCGAGCACCUUCUU-UAUGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG --------......((((...((((((..........((((.....(((....)))..-...))))...........))))))))))......((((....))))...... ( -19.10, z-score = -0.80, R) >droMoj3.scaffold_6540 4336945 110 - 34148556 UAUGCGACUACAUAUGCCACAGCAGGGGCUGCCACAACCACUCGCCAAGCACCUUCUU-UAUGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG ((((......))))((((...((((((((..............))).......((((.-.....))))..........)))))))))......((((....))))...... ( -17.94, z-score = 1.22, R) >consensus __________________ACAUACAUAUAUGUGCUUUUCGCUCUCCUUACCUCUCCGCAAAUGUGGAAAUUUGAAUAUCCUGCGGUAAUUUCCACUAUUUUUAGUACUUUG ..............................................................(((((((((...............)))))))))................ ( -8.86 = -8.86 + 0.00)



| Location | 22,644,629 – 22,644,723 |
|---|---|
| Length | 94 |
| Sequences | 11 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 71.74 |
| Shannon entropy | 0.56126 |
| G+C content | 0.38590 |
| Mean single sequence MFE | -18.18 |
| Consensus MFE | -9.26 |
| Energy contribution | -9.26 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.739727 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
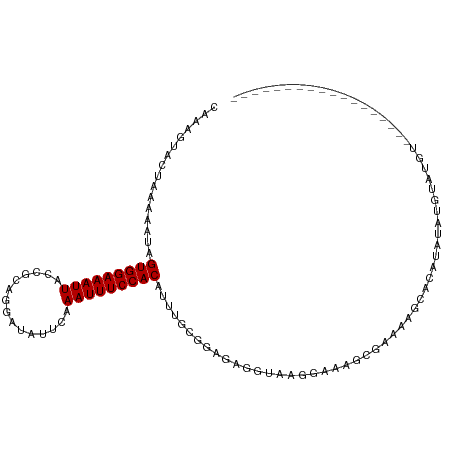
>dm3.chr3R 22644629 94 - 27905053 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACAUUUGCGGAGAGGUAAGGAAAGCGCAAAGCACAUAUAUGUACAUA----------------- ....((((........(((((((((...............))))))))).((((((...............))))))..........))))...----------------- ( -18.22, z-score = -1.88, R) >droSim1.chr3R 22475785 94 - 27517382 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACAUUUGCGGAGAGGUAAGGAAAGCGAAAAGCACAUGCAUAUAUGUA----------------- .....((((.......((((......))))............(((((.(....).)))))))))......((.....))(((((....))))).----------------- ( -14.90, z-score = -0.36, R) >droEre2.scaffold_4820 5071283 94 + 10470090 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACAUUUGCGGAGAGGUAAGAAUAGCGAAAAGCACGUAUACAUAUGUG----------------- ....((((........((((......))))....(((((...(((((.(....).))))).....)))))((.....))..)))).........----------------- ( -16.20, z-score = -1.07, R) >droYak2.chr3R 4984041 94 + 28832112 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACAUUUGCGGAGAGGUAUGGAAAGCGAAAAGUACAUAUAUGUACAUA----------------- ....(((((.......((((......))))......(((...((((((.((((......)))).))))))..))).))))).............----------------- ( -18.10, z-score = -1.85, R) >droSec1.super_13 1418411 94 - 2104621 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACAUUUGCGGAGAGGUAAGGAAAGCGAAAAGCACAUACAUAUAUGUA----------------- .....((((.......((((......))))............(((((.(....).)))))))))......((.....))(((((....))))).----------------- ( -15.20, z-score = -1.07, R) >droWil1.scaffold_181130 13966757 73 + 16660200 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACACACACAAACACACACACCCACAC-------------------------------------- ................(((((((((...............)))))))))........................-------------------------------------- ( -9.26, z-score = -2.54, R) >dp4.chr2 25963407 86 + 30794189 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACA-UUGCGGCGUCG-AAGGUGAGGCCAGAGCAUGUGUGUG----------------------- ................((((......))))................((((-(((((((.(((-....))).)))...))).)))))..----------------------- ( -18.80, z-score = -0.38, R) >droPer1.super_19 1340670 86 + 1869541 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACA-UUGCGGCGUCG-AAGGUGAGGCCAGAGCAUGUGUGUG----------------------- ................((((......))))................((((-(((((((.(((-....))).)))...))).)))))..----------------------- ( -18.80, z-score = -0.38, R) >droAna3.scaffold_13340 7008992 85 + 23697760 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACA-UUGCGGAGAGGUAAGGACAUACAUACGUACAUAAA------------------------- ....((((......((((....))))((((((((..........)))...-.)))))....(((.(......).))))))).....------------------------- ( -15.10, z-score = -1.86, R) >droVir3.scaffold_13047 11963302 102 - 19223366 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACAUA-AAGAAGGUGCUCGGUGAGUGGUUGUGGCAACUCCUGCUGUGGCAUAUGUA-------- ................(((((((((...............)))))))))...-......(((((((((.((..((((...))))..)))))).))))).....-------- ( -22.96, z-score = -0.06, R) >droMoj3.scaffold_6540 4336945 110 + 34148556 CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACAUA-AAGAAGGUGCUUGGCGAGUGGUUGUGGCAGCCCCUGCUGUGGCAUAUGUAGUCGCAUA ..(((((((.......(((((((((...............)))))))))...-.....))))))).((((.((..(((.(((((....))))).)))....)).))))... ( -32.42, z-score = -1.56, R) >consensus CAAAGUACUAAAAAUAGUGGAAAUUACCGCAGGAUAUUCAAAUUUCCACAUUUGCGGAGAGGUAAGGAAAGCGAAAAGCACAUAUAUGUAUGU__________________ ................(((((((((...............))))))))).............................................................. ( -9.26 = -9.26 + -0.00)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:46:42 2011