
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 8,715,726 – 8,715,819 |
| Length | 93 |
| Max. P | 0.896378 |

| Location | 8,715,726 – 8,715,819 |
|---|---|
| Length | 93 |
| Sequences | 8 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 75.84 |
| Shannon entropy | 0.45285 |
| G+C content | 0.42510 |
| Mean single sequence MFE | -22.05 |
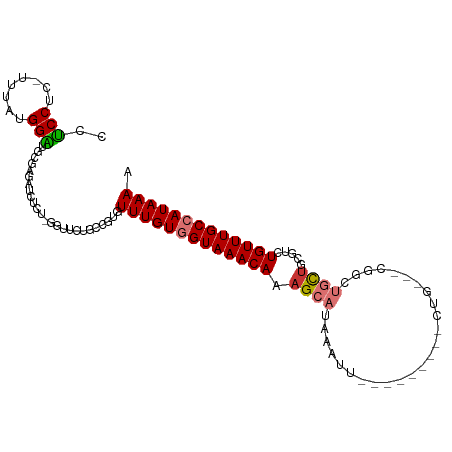
| Consensus MFE | -13.29 |
| Energy contribution | -13.62 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.67 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.896378 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
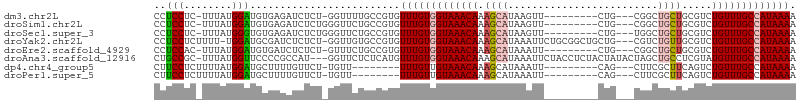
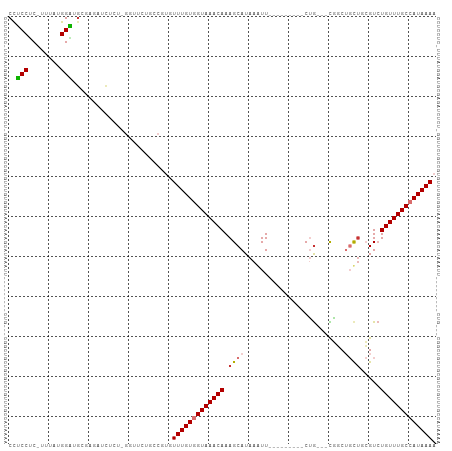
>dm3.chr2L 8715726 93 - 23011544 CCUCCUC-UUUAUGGAUGUGAGAUCUCU-GGUUUUGCCGUGUUUGUGGUAAACAAAGCAUAAGUU---------CUG---CGGCUGCUGCGUCUGUUUGCCAUAAAA ..(((..-.....))).(..((((....-.))))..)....(((((((((((((..(((..(((.---------...---..)))..)))...))))))))))))). ( -26.80, z-score = -2.43, R) >droSim1.chr2L 8500870 94 - 22036055 CCUCCUC-UUUAUGGAUGUGAGAUCUCUGGGUUCUGCCGUGUUUGUGGUAAACAAAGCAUAAGUU---------CUG---CGGCUGCUGCGUCUGUUUGCCAUAAAA .((((((-(....))).).))).......((.....))...(((((((((((((..(((..(((.---------...---..)))..)))...))))))))))))). ( -25.70, z-score = -1.63, R) >droSec1.super_3 4197436 94 - 7220098 CCUCCUC-UUUAUGGGUGUGAGAUCUCUGGGUUCUGCCGUGUUUGUGGUAAACAAAGCAUAAGUU---------CUG---UGGCUGCUGCGUCUGUUUGCCAUAAAA .((((.(-(.....)).).))).......((.....))...(((((((((((((..(((..(((.---------...---..)))..)))...))))))))))))). ( -25.10, z-score = -1.51, R) >droYak2.chr2L 11364373 102 - 22324452 CCUCCUCUUUU-UGGAUGCGAUCUCUCU-GGUUGUGCCGUGUUUGUGGUAAACAAAGCAUAAAUUCUGCGGCUGCUG---CGUCUGUUGCGUCUGUUUGCCAUAAAA ...........-.((.(((((((.....-)))))))))...(((((((((((((..(((.......)))(((.((.(---(....)).)))))))))))))))))). ( -29.60, z-score = -3.03, R) >droEre2.scaffold_4929 9309554 93 - 26641161 CCUCCAC-UUUAUGGAUGUGAUCUCUCU-GUUUCUGCCGUGUUUGUGGUAAACAAAGCAUAAAUU---------CUG---CGGCUGCUGCGUCUGUUUGCCAUAAAA .......-..(((((....(((......-)))....)))))(((((((((((((..(((......---------.))---).((....))...))))))))))))). ( -22.30, z-score = -1.88, R) >droAna3.scaffold_12916 10364034 103 - 16180835 CUGCCGC-UUUAUGGUUCCCCGCCAU---GGUUCUCUCAUGUUUGUGGUAAACAAAGCAUAAAUUCUACCUCUACUAUACUAGCUGCCUCGUAUGUUUGCCAUAAAA .....(.-.(((((((.....)))))---))..).......(((((((((((((.(((........................)))........))))))))))))). ( -19.26, z-score = -0.95, R) >dp4.chr4_group5 1501507 86 + 2436548 CUUCCUCUUUUAUGGAUGCUUUUGUUCU-UGUU--------UUUGUUGUAAACAAAGCAUAAAUU---------CAG---CUUCGCUUCAGUCUGUUUGCCAUAAAA ........((((((((((((((.(((..-((..--------.....))..)))))))))).....---------(((---((.......)).)))....))))))). ( -13.80, z-score = -0.96, R) >droPer1.super_5 1470192 86 + 6813705 CUUCCUCUUUUAUGGAUGCUUUUGUUCU-UGUU--------UUUGUUGUAAACAAAGCAUAAAUU---------CAG---CUUCGCUUCAGUCUGUUUGCCAUAAAA ........((((((((((((((.(((..-((..--------.....))..)))))))))).....---------(((---((.......)).)))....))))))). ( -13.80, z-score = -0.96, R) >consensus CCUCCUC_UUUAUGGAUGCGAGAUCUCU_GGUUCUGCCGUGUUUGUGGUAAACAAAGCAUAAAUU_________CUG___CGGCUGCUGCGUCUGUUUGCCAUAAAA ..(((........))).........................(((((((((((((.((((.........................)))).....))))))))))))). (-13.29 = -13.62 + 0.33)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:26:42 2011