
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 21,917,671 – 21,917,764 |
| Length | 93 |
| Max. P | 0.677382 |

| Location | 21,917,671 – 21,917,764 |
|---|---|
| Length | 93 |
| Sequences | 8 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 70.79 |
| Shannon entropy | 0.54227 |
| G+C content | 0.48448 |
| Mean single sequence MFE | -22.50 |
| Consensus MFE | -11.22 |
| Energy contribution | -11.44 |
| Covariance contribution | 0.22 |
| Combinations/Pair | 1.55 |
| Mean z-score | -1.24 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.677382 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
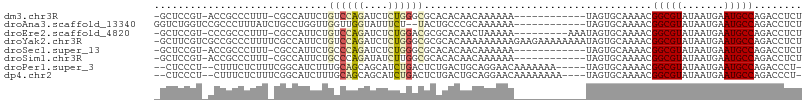
>dm3.chr3R 21917671 93 + 27905053 -GCUCCGU-ACCGCCCUUU-CGCCAUUCUGUCCAGAUCUCUGGGCGCACACAACAAAAAA------------UAGUGCAAAACGGCGUAUAAUGAAUGCCAGACCUCU -.....((-(.........-((((.....((((((....))))))((((...........------------..)))).....)))).........)))......... ( -22.09, z-score = -1.50, R) >droAna3.scaffold_13340 15743914 94 + 23697760 GGUCUGGUCCGCCCUUUAUCUGCCUGGUUGGUUGGUAUUUCU--UACUGCCCGCAAAAAA------------UAGUGCAAAACGGCGUAUAAUGAAUGCCAGACCUCU ((((((((...(...((((.((((.....((..((((.....--))))..))(((.....------------...))).....)))).)))).)...))))))))... ( -29.30, z-score = -2.55, R) >droEre2.scaffold_4820 4344376 96 - 10470090 -GCUCCGU-CCCGCCCUUU-CGCCAUUCUGUCCAGAUCUCUGGACGCGCACAACUAAAAA---------AAAUAGUGCAAAACGGCGUAUAAUGAAUGCCAGACCUCU -.....((-(..((..((.-((((.....((((((....))))))..((((.........---------.....)))).....))))......))..))..))).... ( -24.24, z-score = -2.87, R) >droYak2.chr3R 4275758 107 - 28832112 -GCUUCGUCGCCGCCCUUUUCGCCAUUCUGUCCAGAUCUCUGGGCGCGCACAAAAAAAAAGAAGAAAAAAAAUAGUGCAAAACGGCGUAUAAUGAAUGCCAGACCUCU -((((((((((((................((((((....))))))..((((.......................))))....)))))....))))).))......... ( -25.80, z-score = -1.28, R) >droSec1.super_13 712169 93 + 2104621 -GCUCCGU-ACCGCCCUUU-CGCCAUUCUGCCCAGAUCUCUGGGCGCACACAACAAAAAA------------UAGUGCAAAACGGCGUAUAAUGAAUGCCAGACCUCU -.....((-(.........-((((.....((((((....))))))((((...........------------..)))).....)))).........)))......... ( -24.79, z-score = -2.32, R) >droSim1.chr3R 21778220 93 + 27517382 -GCUCCGU-ACCGCCCUUU-CGCCAUUCUGCCCAGAUAUCUUGGCGCACACAACAAAAAA------------UAGUGCAAAACGGCGUAUAAUGAAUGCCAGACCUCU -((..(((-..((((....-.((((.(((....))).....))))((((...........------------..)))).....))))....)))...))......... ( -18.62, z-score = -0.72, R) >droPer1.super_3 5511200 98 - 7375914 --CUCCCU--CUUUCUCUUUCGGCAUCUUUGCAGCAGCAUCUGACUCUGACUGCAGGAACAAAAAAA-----UAGUGCAAAACGGCGUAUAAUGAAUGCCAGACCCU- --......--...........(((((.((((((((((.........))).)))))))..........-----..((((........)))).....))))).......- ( -17.60, z-score = 0.65, R) >dp4.chr2 11674688 99 - 30794189 --CUCCCU--CUUUCUCUUUCGGCAUCUUUGCAGCAGCAUCUGACUCUGACUGCAGGAACAAAAAAAA----UAGUGCAAAACGGCGUAUAAUGAAUGCCAGACCCU- --......--...........(((((.((((((((((.........))).)))))))...........----..((((........)))).....))))).......- ( -17.60, z-score = 0.65, R) >consensus _GCUCCGU_ACCGCCCUUU_CGCCAUUCUGGCCAGAUCUCUGGGCGCACACAACAAAAAA____________UAGUGCAAAACGGCGUAUAAUGAAUGCCAGACCUCU .............................((((((....))))))......................................(((((.......)))))........ (-11.22 = -11.44 + 0.22)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:44:46 2011