| Sequence ID | dm3.chr3R |
|---|---|
| Location | 21,846,252 – 21,846,343 |
| Length | 91 |
| Max. P | 0.952824 |

| Location | 21,846,252 – 21,846,343 |
|---|---|
| Length | 91 |
| Sequences | 10 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 73.13 |
| Shannon entropy | 0.49270 |
| G+C content | 0.52460 |
| Mean single sequence MFE | -23.03 |
| Consensus MFE | -12.26 |
| Energy contribution | -12.23 |
| Covariance contribution | -0.03 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.59 |
| SVM RNA-class probability | 0.952824 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
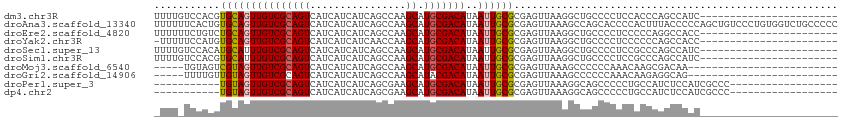
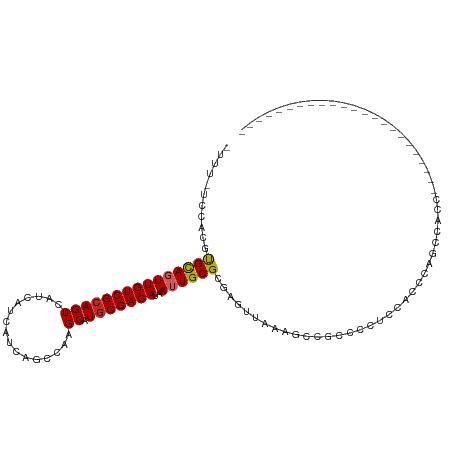
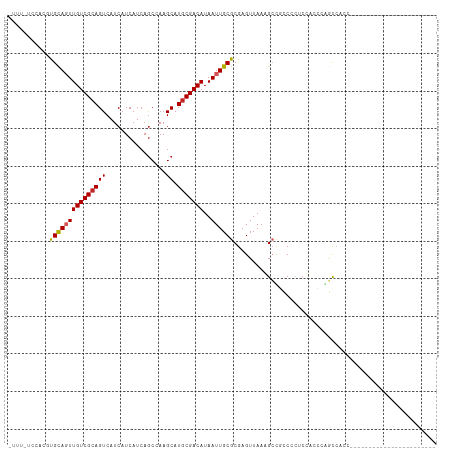
>dm3.chr3R 21846252 91 + 27905053 UUUUGUCCACGUGCAGUUGUCGCAGUCAUCAUCAUCAGCCAAGCAUGCGACAUAAUUGCGCGAGUUAAGGCUGCCCCUCCACCCAGCCAUC----------------------- .........(((((((((((((((((................)).)))))))..))))))))......(((((..........)))))...----------------------- ( -27.19, z-score = -3.20, R) >droAna3.scaffold_13340 12399441 114 - 23697760 UUUUUUCACUGUGCAGUUGUCGCAGUCAUCAUCAUCAGCCAAGCAUGCGACAUAAUUGCGCGAGUUAAAGCCAGCACCCCACUUUACCCCCAGCUGUCCCUGUGGUCUGCCCCC ..((((.(((((((((((((((((((................)).)))))))..))))))).))).))))...(((..((((...................))))..))).... ( -28.00, z-score = -1.97, R) >droEre2.scaffold_4820 4269826 91 - 10470090 UUUUUUCUGUCUGCAGUUGUCGCAGUCAUCAUCAUCAGCCAAGCAUGCGACAUAAUUGCGCGAGUUAAGGCUGCCCCUCCCCCAGGCCACC----------------------- .......((.((((.......)))).)).......(((((.(((.((((.((....)))))).)))..)))))..(((.....))).....----------------------- ( -21.40, z-score = -0.71, R) >droYak2.chr3R 4202412 90 - 28832112 -UUUUUCCAUGUGCAGUUGUCGCAGUCAUCAUCAUCAACCAAGCAUGCGACAUAAUUGCGCGAGUUAAGGCUGCCCCUCCCCCCAGCCACC----------------------- -........(((((((((((((((((................)).)))))))..))))))))......(((((..........)))))...----------------------- ( -25.19, z-score = -3.72, R) >droSec1.super_13 640772 91 + 2104621 UUUUGUCCACAUGCAUUUGUCGCAGUCAUCAUCAUCAGCCAAGCAUGCGACAUAAUUGCGCGAGUUAAGGCUGCCCCUCCGCCCAGCCAUC----------------------- .........(.((((..(((((((((................)).)))))))....)))).)......(((((..(....)..)))))...----------------------- ( -20.09, z-score = -0.53, R) >droSim1.chr3R 21697387 91 + 27517382 UUUUGUCCACGUGCAUUUGUCGCAGUCAUCAUCAUCAGCCAAGCAUGCGACAUAAUUGCGCGAGUUAAGGCUGCCCCUCCGCCCAGCCAUC----------------------- .........((((((..(((((((((................)).)))))))....))))))......(((((..(....)..)))))...----------------------- ( -26.09, z-score = -2.48, R) >droMoj3.scaffold_6540 15798482 84 + 34148556 -----UGUAGUCGUAGUUGUCGCAGUCAUCAUCAUCAGCCAAGCAUGCGACAUAAUUGCGCGAGUUAAAGCCCCCCAAACAAGCGACAA------------------------- -----....(((((((((((((((((................)).)))))))..)))))((..(((...........)))..)))))..------------------------- ( -18.29, z-score = -1.69, R) >droGri2.scaffold_14906 10803362 84 + 14172833 -----UUUUGUUGUAGUUGUCGCAGUCAUCAUCAUCAGCCAAGCAUACGACAUAAUUGCGCGAGUUAAAGCCCCCCAAACAAGAGGCAG------------------------- -----...((((((((((...((..............))..))).))))))).....((....))....(((.(........).)))..------------------------- ( -13.44, z-score = -0.07, R) >droPer1.super_3 5428227 85 - 7375914 -----------UGUAGUUGUCGCAGUCAUCAUCAUCAGCGAAGCAUGCGACAUAAUUGCGCGAGUUAAAGGCAGCCCCCUGCCAUCUCCAUCGCCC------------------ -----------.((((((((((((....((.........))....)))))))..)))))((((......(((((....))))).......))))..------------------ ( -25.32, z-score = -2.66, R) >dp4.chr2 11592358 85 - 30794189 -----------UGUAGUUGUCGCAGUCAUCAUCAUCAGCGAAGCAUGCGACAUAAUUGCGCGAGUUAAAGGCAGCCCCCUGCCAUCUCCAUCGCCC------------------ -----------.((((((((((((....((.........))....)))))))..)))))((((......(((((....))))).......))))..------------------ ( -25.32, z-score = -2.66, R) >consensus _UUU_UCCACGUGCAGUUGUCGCAGUCAUCAUCAUCAGCCAAGCAUGCGACAUAAUUGCGCGAGUUAAAGCCGCCCCUCCACCCAGCCACC_______________________ ...........(((((((((((((((................)).)))))))..))))))...................................................... (-12.26 = -12.23 + -0.03)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:44:31 2011