| Sequence ID | dm3.chr3R |
|---|---|
| Location | 21,728,515 – 21,728,569 |
| Length | 54 |
| Max. P | 0.961589 |

| Location | 21,728,515 – 21,728,569 |
|---|---|
| Length | 54 |
| Sequences | 12 |
| Columns | 61 |
| Reading direction | forward |
| Mean pairwise identity | 68.04 |
| Shannon entropy | 0.65009 |
| G+C content | 0.47980 |
| Mean single sequence MFE | -12.35 |
| Consensus MFE | -6.30 |
| Energy contribution | -6.57 |
| Covariance contribution | 0.26 |
| Combinations/Pair | 1.67 |
| Mean z-score | -1.35 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.70 |
| SVM RNA-class probability | 0.961589 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
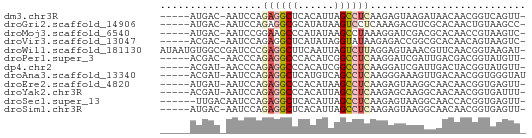
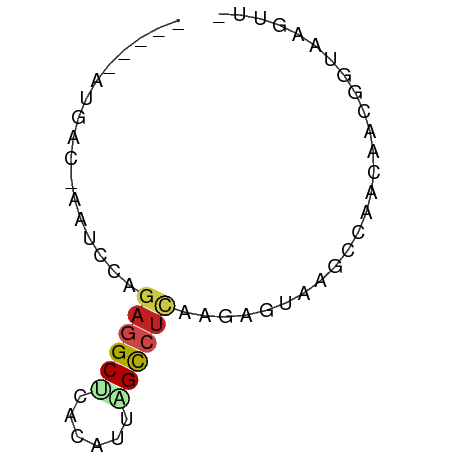
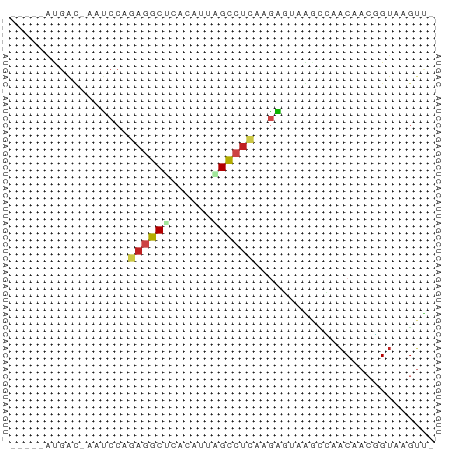
>dm3.chr3R 21728515 54 + 27905053 -----AUGAC-AAUCCAGAGGCUCACAUUAGCCUCAAGAGUAAGAUAACAACGGUCAGUU- -----.((((-......((((((......))))))....((......))....))))...- ( -13.40, z-score = -3.02, R) >droGri2.scaffold_14906 10672673 54 + 14172833 -----AUGAC-AAUCCAGAGGCGCAUAUAAGUCCUCAAAGACGUCGCACAACUGUAAGCC- -----..(((-......((((.((......))))))......)))...............- ( -7.70, z-score = -0.01, R) >droMoj3.scaffold_6540 15642798 54 + 34148556 -----AUGAC-AAUCCGGAAGCCCAUAUAAGCCUAAAGGAUCGACGCACAACCGUAAGUC- -----..(((-.((((....((........)).....))))..(((......)))..)))- ( -6.10, z-score = 0.46, R) >droVir3.scaffold_13047 5279589 54 + 19223366 -----ACGAC-AAUCCAGAGGCUCAUAUAGGUAUAAGAGACCGGCGCACAACAGUAAGUC- -----..(((-........(((((.(((....))).))).)).((........))..)))- ( -7.40, z-score = -0.10, R) >droWil1.scaffold_181130 2370491 60 - 16660200 AUAAUGUGGCCGAUCCCGAGGCUUCAAUUAGUCUUAGGAGUAAACGUUCAACGGUAAGAU- ........((((.(((..(((((......)))))..)))(....)......)))).....- ( -14.80, z-score = -1.40, R) >droPer1.super_3 5297073 54 - 7375914 -----ACGAC-AACCCAGAGGCCCACAUCGGCCUCAAGGAUCGAUUGACGACGGUAUGUU- -----..(((-(..((.((((((......)))))).....(((.....))).))..))))- ( -17.10, z-score = -1.99, R) >dp4.chr2 11461801 54 - 30794189 -----ACGAU-AACCCAGAGGCCCACAUCGGCCUCAAGGAUCGAUUGACUACGGUAUGUU- -----.((((-..((..((((((......))))))..))))))....((....)).....- ( -17.80, z-score = -2.59, R) >droAna3.scaffold_13340 12276608 55 - 23697760 -----ACGAU-AAUCCAGAGGCUCAUGUCAGCCUCAAGGGAAAGUUGACAACGGUGGGUAU -----.((((-..(((.((((((......))))))...)))..)))).............. ( -15.00, z-score = -1.39, R) >droEre2.scaffold_4820 4149675 54 - 10470090 -----AUGAU-AAUCCAGAGGCCCACAUAAGCCUCAAGAGUAAGGCAACAACGGUGAGUU- -----.....-...((.(((((........)))))........(....)...))......- ( -11.10, z-score = -1.25, R) >droYak2.chr3R 4079255 54 - 28832112 -----ACGAU-AAUCCAGAGGCCCACAUUAGCCUCAAGAGCAAGGCAACAACGGUGAUUU- -----..(((-...((.(((((........)))))........(....)...))..))).- ( -11.50, z-score = -1.65, R) >droSec1.super_13 511932 54 + 2104621 ------UUGACAAUCCAGAGGCUCACAUUAGCCUCAAGAGUAAGGCAACCACGGUGAGUU- ------..(((...((.((((((......))))))........(....)...))...)))- ( -12.70, z-score = -1.34, R) >droSim1.chr3R 21573176 54 + 27517382 -----AUGAC-AAUCCAGAGGCUCACAUUAGCCUCAAGAGUAAGGCAACAACGGUGAGUU- -----..(((-...((.((((((......))))))........(....)...))...)))- ( -13.60, z-score = -1.97, R) >consensus _____AUGAC_AAUCCAGAGGCUCACAUUAGCCUCAAGAGUAAGCCAACAACGGUAAGUU_ .................((((((......)))))).......................... ( -6.30 = -6.57 + 0.26)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:44:22 2011