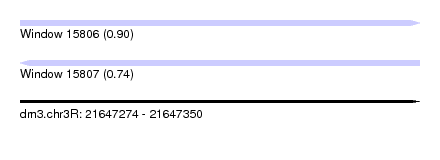

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 21,647,274 – 21,647,350 |
| Length | 76 |
| Max. P | 0.895392 |

| Location | 21,647,274 – 21,647,350 |
|---|---|
| Length | 76 |
| Sequences | 9 |
| Columns | 77 |
| Reading direction | forward |
| Mean pairwise identity | 66.25 |
| Shannon entropy | 0.72768 |
| G+C content | 0.49745 |
| Mean single sequence MFE | -22.72 |
| Consensus MFE | -6.29 |
| Energy contribution | -6.72 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.40 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.28 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.12 |
| SVM RNA-class probability | 0.895392 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 21647274 76 + 27905053 UGGCGGAAUGCCAAACGAUGCCGAAAGGUCGCAGAAAUGGCAGCCAUCUAAUGGCAUCGAGACAUUUAAAAA-CAUU ((((.....))))..((((((((..((((.((..........)).))))..)))))))).............-.... ( -21.80, z-score = -2.10, R) >droSim1.chr3R 21477052 76 + 27517382 UGGCGGAAUGCCAAAGGAUGCCGAAAGGUCGCAGGAAUGGCAGCCACCUAAUGGCAUCGAGCCAUUUAAAAA-CAUC ((((((....))....(((((((..((((.((..........)).))))..)))))))..))))........-.... ( -24.70, z-score = -2.01, R) >droSec1.super_13 436097 75 + 2104621 UGGCGGAAUGCCAAAGGAUGCCGAA-GGUCGCAGGAAUGGCAGCCAUCUAAUGGCAUCGAGCCAUUUAAAAA-CAUC ((((...((((((..(((((.(...-.((((......)))).).)))))..))))))...))))........-.... ( -22.80, z-score = -1.42, R) >droYak2.chr3R 4006069 76 - 28832112 UGGCGGAAUGCCAAAGGAUGCCGAAAGCUGGCAGGAAUGGCAGCCAUCUAAUGGCAUCGAGCCAUUUAAAAA-CAUC ((((...((((((..(((((......((((.((....)).)))))))))..))))))...))))........-.... ( -27.10, z-score = -2.66, R) >droEre2.scaffold_4820 4077034 76 - 10470090 UGGCGGAAUGCCAAAGGAUGCCGAAAGGUUGCAGGAAUGGCAGCCAUCUAAUGGCGUCGAGCCAUUUAAAAA-CAUC ((((...((((((..(((((.(....)(((((.......))))))))))..))))))...))))........-.... ( -26.00, z-score = -2.43, R) >droAna3.scaffold_13340 12204959 60 - 23697760 -----------------UGGCCACCAGCCACUUGGAAUGCCACCAGUCUUGUGGCAUCGCGCCAUUUAAAAAACAUC -----------------(((((.((((....)))).(((((((.......))))))).).))))............. ( -17.00, z-score = -1.71, R) >dp4.chr2 7353952 62 - 30794189 UGGCGGAAUGCCAC----UGCCAU----CCGCAGGAUGAG-----AUCUAAUGGCAUCGAGCCAUUUAAAAA-CAU- ((((...((((((.----..((((----((...))))).)-----......))))))...))))........-...- ( -18.50, z-score = -1.74, R) >droPer1.super_3 1164273 63 - 7375914 UGGCGGAAUGCCAU----UGCCAU----CCGCAGGAUGAG-----AUCUAAUGGCAUCGAGCCAUUUAAAAA-CAUC ((((...(((((((----((((((----((...))))).)-----...)))))))))...))))........-.... ( -21.30, z-score = -2.61, R) >droWil1.scaffold_181130 2265362 75 - 16660200 GGGCCACAAGACAGACGGGAACUUGAUGUGACACAGACAC--ACAACCUGGUGUUGUCUUUUUGGCCAAGAAGUGGC .(((((.(((((((.((((...(((.((((.......)))--))))))))...)))))))..))))).......... ( -25.30, z-score = -1.71, R) >consensus UGGCGGAAUGCCAAAGGAUGCCGAAAGGUCGCAGGAAUGGCAGCCAUCUAAUGGCAUCGAGCCAUUUAAAAA_CAUC ((((.....))))....................................((((((.....))))))........... ( -6.29 = -6.72 + 0.44)


| Location | 21,647,274 – 21,647,350 |
|---|---|
| Length | 76 |
| Sequences | 9 |
| Columns | 77 |
| Reading direction | reverse |
| Mean pairwise identity | 66.25 |
| Shannon entropy | 0.72768 |
| G+C content | 0.49745 |
| Mean single sequence MFE | -21.71 |
| Consensus MFE | -6.67 |
| Energy contribution | -7.02 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.57 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.740541 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 21647274 76 - 27905053 AAUG-UUUUUAAAUGUCUCGAUGCCAUUAGAUGGCUGCCAUUUCUGCGACCUUUCGGCAUCGUUUGGCAUUCCGCCA ....-......((((((.(((((((...((.(.((..........)).).))...)))))))...))))))...... ( -20.30, z-score = -1.49, R) >droSim1.chr3R 21477052 76 - 27517382 GAUG-UUUUUAAAUGGCUCGAUGCCAUUAGGUGGCUGCCAUUCCUGCGACCUUUCGGCAUCCUUUGGCAUUCCGCCA ....-......((((((.....)))))).(((((.(((((....(((((....)).))).....)))))..))))). ( -25.50, z-score = -2.11, R) >droSec1.super_13 436097 75 - 2104621 GAUG-UUUUUAAAUGGCUCGAUGCCAUUAGAUGGCUGCCAUUCCUGCGACC-UUCGGCAUCCUUUGGCAUUCCGCCA ((((-((....((((((.....)))))).((.((.(((.......))).))-.))))))))...((((.....)))) ( -21.90, z-score = -1.36, R) >droYak2.chr3R 4006069 76 + 28832112 GAUG-UUUUUAAAUGGCUCGAUGCCAUUAGAUGGCUGCCAUUCCUGCCAGCUUUCGGCAUCCUUUGGCAUUCCGCCA ((((-((....((((((.....)))))).((.(((((.((....)).))))).))))))))...((((.....)))) ( -25.50, z-score = -2.21, R) >droEre2.scaffold_4820 4077034 76 + 10470090 GAUG-UUUUUAAAUGGCUCGACGCCAUUAGAUGGCUGCCAUUCCUGCAACCUUUCGGCAUCCUUUGGCAUUCCGCCA ((((-((....((((((.....)))))).((.((.(((.......))).))..))))))))...((((.....)))) ( -21.10, z-score = -1.48, R) >droAna3.scaffold_13340 12204959 60 + 23697760 GAUGUUUUUUAAAUGGCGCGAUGCCACAAGACUGGUGGCAUUCCAAGUGGCUGGUGGCCA----------------- .............(((...((((((((.......)))))))))))..((((.....))))----------------- ( -18.00, z-score = -0.30, R) >dp4.chr2 7353952 62 + 30794189 -AUG-UUUUUAAAUGGCUCGAUGCCAUUAGAU-----CUCAUCCUGCGG----AUGGCA----GUGGCAUUCCGCCA -...-........((((..(((((((((.(..-----.((((((...))----))))))----))))))))..)))) ( -21.10, z-score = -1.97, R) >droPer1.super_3 1164273 63 + 7375914 GAUG-UUUUUAAAUGGCUCGAUGCCAUUAGAU-----CUCAUCCUGCGG----AUGGCA----AUGGCAUUCCGCCA ((((-...((.((((((.....)))))).)).-----..))))..((((----(..((.----...))..))))).. ( -21.60, z-score = -2.10, R) >droWil1.scaffold_181130 2265362 75 + 16660200 GCCACUUCUUGGCCAAAAAGACAACACCAGGUUGU--GUGUCUGUGUCACAUCAAGUUCCCGUCUGUCUUGUGGCCC ..........(((((..((((((((...((.((((--(((.......)))).))).))...)).)))))).))))). ( -20.40, z-score = -1.10, R) >consensus GAUG_UUUUUAAAUGGCUCGAUGCCAUUAGAUGGCUGCCAUUCCUGCGACCUUUCGGCAUCCUUUGGCAUUCCGCCA .......(((((.((((.....)))))))))..................................(((.....))). ( -6.67 = -7.02 + 0.36)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:44:10 2011