
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 21,541,473 – 21,541,564 |
| Length | 91 |
| Max. P | 0.633369 |

| Location | 21,541,473 – 21,541,564 |
|---|---|
| Length | 91 |
| Sequences | 11 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 66.74 |
| Shannon entropy | 0.66673 |
| G+C content | 0.51155 |
| Mean single sequence MFE | -15.98 |
| Consensus MFE | -10.01 |
| Energy contribution | -9.70 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.20 |
| Mean z-score | -0.12 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.633369 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
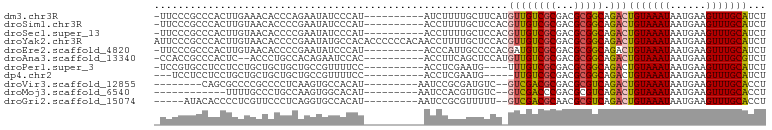
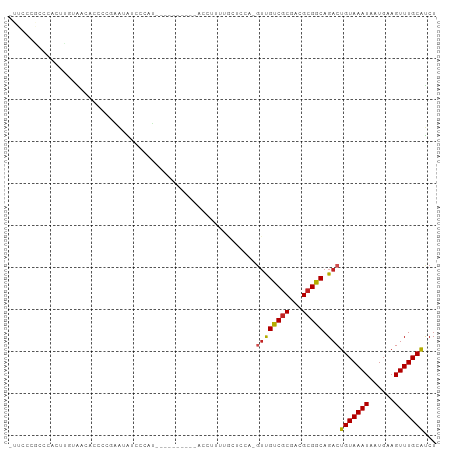
>dm3.chr3R 21541473 91 - 27905053 -UUCCCGCCCACUUGAAACACCCAGAAUAUCCCAU----------AUCUUUUGCUUCAUGUUGUCGCGACGCGGCAGACUGUAAAUAAUGAAGUUUGCAUCU -....(((.....((((.((...(((.........----------.)))..)).)))).((((...)))))))(((((((...........))))))).... ( -14.30, z-score = 0.15, R) >droSim1.chr3R 21377198 91 - 27517382 -UUCCCGCCCACUUGUAACACCCCGAAUAUCCCAU----------ACCUUUUGCUCCACGUUGUCGCGACGCGGCAGACUGUAAAUAAUGAAGUUUGCAUCU -....(((....(((........))).........----------.....((((..(.....)..)))).)))(((((((...........))))))).... ( -12.90, z-score = 0.38, R) >droSec1.super_13 338823 91 - 2104621 -UUCCCGCCCACUUGUAACACCCCGAAUAUCCCAU----------ACCUUUUGCUCCACGUUGUCGCGACGCGGCAGACUGUAAAUAAUGAAGUUUGCAUCU -....(((....(((........))).........----------.....((((..(.....)..)))).)))(((((((...........))))))).... ( -12.90, z-score = 0.38, R) >droYak2.chr3R 3904255 102 + 28832112 AUUCCCGCCCACUUGUAACACCCCGAAUAUGCCACACCCCCCACAACCUUUUGCUCCACGUUGUCGCGACGCGGCAGACUGUAAAUAAUGAAGUUUGCAUCU .....(((....(((..(((...((.....((....................))....)).)))..))).)))(((((((...........))))))).... ( -13.85, z-score = 0.53, R) >droEre2.scaffold_4820 3972704 91 + 10470090 -UUCCCGCCCACUUGUAACACCCCGAAUAUCCCAU----------ACCCAUUGCCCCACGAUGUCGCGACGCGGCAGACUGUAAAUAAUGAAGUUUGCAUCU -....(((....(((........))).........----------...(((((.....))))).......)))(((((((...........))))))).... ( -12.90, z-score = 0.14, R) >droAna3.scaffold_13340 12108561 89 + 23697760 -CCACCGCCCACUC--ACCCUGCCACAGAAUCCAC----------ACCUUCAGCUCCAUGUUGUCGCGACGCGGCAGACUGUAAAUAAUGAAGUUUGCGUCU -...((((......--...(((...))).......----------.....((((.....)))).......)))).((((.((((((......)))))))))) ( -15.60, z-score = 0.46, R) >droPer1.super_3 1050785 87 + 7375914 -UCCGUGCCUCCUCCUGCUGCUGCUGCCGUUUUCC----------ACCUCGAAUG----UUUGUCGCGACGCGGCAGACUGUAAAUAAUGAAGUUUGCAUCU -.............((((((((((.(((((((...----------.....)))))----...)).)))..)))))))..(((((((......)))))))... ( -19.20, z-score = -0.17, R) >dp4.chr2 7240981 84 + 30794189 ---UCCUCCUCCUGCUGCUGCUGCUGCCGUUUUCC----------ACCUCGAAUG-----UUGUCGCGACGCGGCAGACUGUAAAUAAUGAAGUUUGCAUCU ---........((((((((((.((...(((((...----------.....)))))-----..)).)))..)))))))..(((((((......)))))))... ( -20.80, z-score = -1.16, R) >droVir3.scaffold_12855 6971180 83 - 10161210 --------CAGCGCCCCGCCCCUCAAGUGCCACAU---------AAUCCGCGAUGUC--GUCGACGCGACGCGUCAGACUGUAAAUAAUGAAGUUUGCACCU --------..(((...))).......((((.....---------.......((((.(--((((...)))))))))(((((...........))))))))).. ( -18.00, z-score = -0.17, R) >droMoj3.scaffold_6540 8503022 79 + 34148556 ------------UUUUGCCCUGCCAAGUGGCACAU---------AAUCCACGUUGUC--GUCGACCCGACGCGUCAGACUGUAAAUAAUGAAGUUUGCACCU ------------...((((((....)).))))...---------.....((((.(((--(......)))))))).....(((((((......)))))))... ( -17.60, z-score = -0.70, R) >droGri2.scaffold_15074 1910161 86 + 7742996 -----AUACACCCCUCGUUCCCUCAGGUGCCACAU---------AAUCCGCGUUUUU--GUCGACGCAACGCGUCAGACUGUAAAUAAUGAAGUUUGCACCU -----...................((((((.....---------..((...((((..--((((((((...))))).)))...))))...)).....)))))) ( -17.70, z-score = -1.09, R) >consensus _UUCCCGCCCACUUGUAACACCCCGAAUAUCCCAU__________ACCUUUUGCUCCA_GUUGUCGCGACGCGGCAGACUGUAAAUAAUGAAGUUUGCAUCU ...........................................................((((((((...))))).)))(((((((......)))))))... (-10.01 = -9.70 + -0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:43:54 2011