| Sequence ID | dm3.chr3R |
|---|---|
| Location | 21,064,795 – 21,064,850 |
| Length | 55 |
| Max. P | 0.987329 |

| Location | 21,064,795 – 21,064,850 |
|---|---|
| Length | 55 |
| Sequences | 5 |
| Columns | 59 |
| Reading direction | forward |
| Mean pairwise identity | 66.26 |
| Shannon entropy | 0.55502 |
| G+C content | 0.38307 |
| Mean single sequence MFE | -13.80 |
| Consensus MFE | -8.40 |
| Energy contribution | -9.44 |
| Covariance contribution | 1.04 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.27 |
| SVM RNA-class probability | 0.987329 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
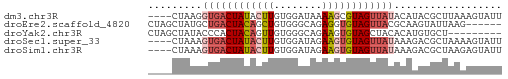
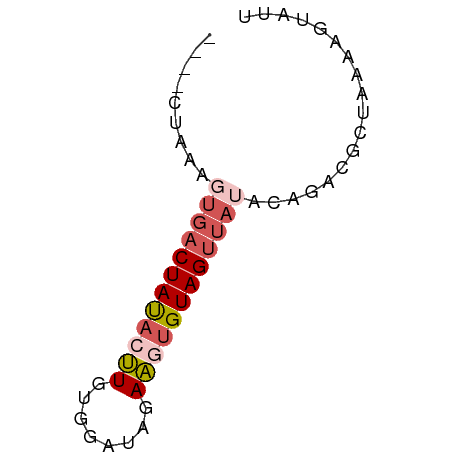
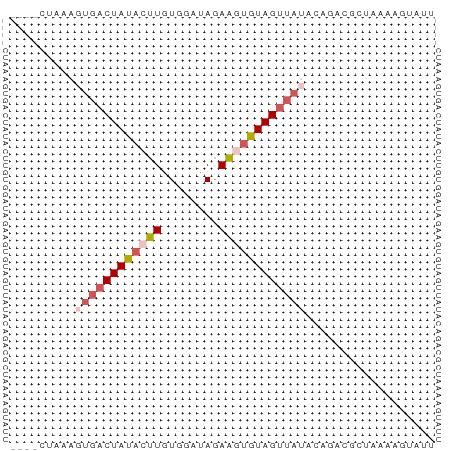
>dm3.chr3R 21064795 55 + 27905053 ----CUAAGGUGACUAUACUUGUGGAUAAAAGCGUAGUUAUACAUACGCUUAAAGUAUU ----...((....))((((((........(((((((........))))))).)))))). ( -11.90, z-score = -1.65, R) >droEre2.scaffold_4820 3491670 53 - 10470090 CUAGCUAUGCUGACUACAGCUGUGGGCAGAGGUGUAGUUACGCAAGUAUUAAG------ ...(((.(((((((((((.((.(....).)).)))))))).))))))......------ ( -17.30, z-score = -1.48, R) >droYak2.chr3R 3406502 50 - 28832112 CUAGCUAUACCCACUACAGUUGUGGGCAGAAGUGUAGCUACACAUGUGCU--------- .(((((((((((((.......)))))......))))))))..........--------- ( -14.80, z-score = -0.40, R) >droSec1.super_33 325392 55 + 478481 ----CUAAAGUGACUAUACUUGUGGAUAGAAGUGUAGUUAUAAAGACGCUAAAAGUAUU ----.....((((((((((((........)))))))))))).................. ( -12.50, z-score = -2.76, R) >droSim1.chr3R 20889815 55 + 27517382 ----CUAAAGUGACUAUACUUGUGGAUAGAAGUGUAGUUAUAAAGACGCUAAGAGUAUU ----.....((((((((((((........)))))))))))).................. ( -12.50, z-score = -2.47, R) >consensus ____CUAAAGUGACUAUACUUGUGGAUAGAAGUGUAGUUAUACAGACGCUAAAAGUAUU .........((((((((((((........)))))))))))).................. ( -8.40 = -9.44 + 1.04)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:42:55 2011