| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,922,219 – 20,922,321 |
| Length | 102 |
| Max. P | 0.863458 |

| Location | 20,922,219 – 20,922,321 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 74.16 |
| Shannon entropy | 0.35272 |
| G+C content | 0.42289 |
| Mean single sequence MFE | -12.50 |
| Consensus MFE | -9.56 |
| Energy contribution | -9.57 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.76 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.622452 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 20922219 102 + 27905053 UGCUUUUGUUUCCAUUAGCUGUCAAUCAUCCUUACAUAUGUUUGACUUCAACCCUAGUCGGAACACCUCGCCUUCGCUACCAUCCAUAUCCAUAUCCACAUC .((...((((((.(((((..(((((.(((........))).))))).......))))).))))))....))............................... ( -13.10, z-score = -1.69, R) >droAna3.scaffold_13340 6783910 85 + 23697760 AGCCGUUGUUUCCAUUAGCUGCCAAUCAUCCUUAUAUAUGUUUGACUUCGACCCUAGUUGGAAUGCCUCGUGUAAGUAUGUAUUC----------------- .((.((((.......)))).))....(((.(((((((...((..(((........)))..)).......))))))).))).....----------------- ( -13.10, z-score = -0.20, R) >droEre2.scaffold_4820 3341860 94 - 10470090 UGCUUUUGUUUCCAUUAGCUGUCAAUCAUCCUUACAUAUGUUUGACUUCAACCCUAUUCGGAACACCUCGCUUUCGCCUCCAUCCAUAUCCACA-------- .(((..((....))..))).(((((.(((........))).))))).............(((.......((....)).....))).........-------- ( -11.30, z-score = -1.65, R) >consensus UGCUUUUGUUUCCAUUAGCUGUCAAUCAUCCUUACAUAUGUUUGACUUCAACCCUAGUCGGAACACCUCGCCUUCGCAUCCAUCCAUAUCCA_A________ .((...((((((.(((((..(((((.(((........))).))))).......))))).))))))....))............................... ( -9.56 = -9.57 + 0.01)


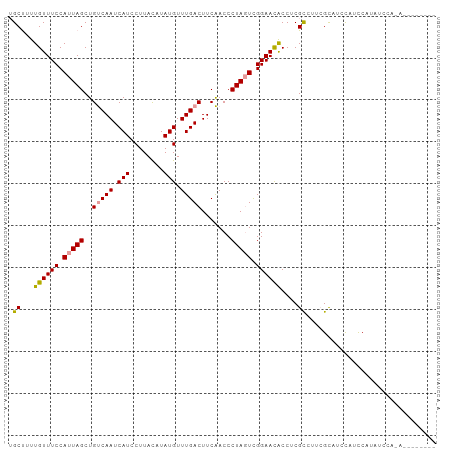
| Location | 20,922,219 – 20,922,321 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 74.16 |
| Shannon entropy | 0.35272 |
| G+C content | 0.42289 |
| Mean single sequence MFE | -18.80 |
| Consensus MFE | -14.77 |
| Energy contribution | -15.00 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.79 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.863458 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 20922219 102 - 27905053 GAUGUGGAUAUGGAUAUGGAUGGUAGCGAAGGCGAGGUGUUCCGACUAGGGUUGAAGUCAAACAUAUGUAAGGAUGAUUGACAGCUAAUGGAAACAAAAGCA ..(((.(((.((.(((((..(((..((....)).........((((....))))...)))..))))).))......))).)))(((..((....))..))). ( -21.70, z-score = -1.40, R) >droAna3.scaffold_13340 6783910 85 - 23697760 -----------------GAAUACAUACUUACACGAGGCAUUCCAACUAGGGUCGAAGUCAAACAUAUAUAAGGAUGAUUGGCAGCUAAUGGAAACAACGGCU -----------------...............(((.....(((.....))))))..(((((.(((........))).)))))((((..((....))..)))) ( -14.60, z-score = -0.98, R) >droEre2.scaffold_4820 3341860 94 + 10470090 --------UGUGGAUAUGGAUGGAGGCGAAAGCGAGGUGUUCCGAAUAGGGUUGAAGUCAAACAUAUGUAAGGAUGAUUGACAGCUAAUGGAAACAAAAGCA --------........((((.....((....)).......))))............(((((.(((........))).))))).(((..((....))..))). ( -20.10, z-score = -2.01, R) >consensus ________U_UGGAUAUGGAUGGAAGCGAAAGCGAGGUGUUCCGACUAGGGUUGAAGUCAAACAUAUGUAAGGAUGAUUGACAGCUAAUGGAAACAAAAGCA .........................((....)).......(((.....))).....(((((.(((........))).))))).(((..((....))..))). (-14.77 = -15.00 + 0.23)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:42:30 2011