
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 797,274 – 797,372 |
| Length | 98 |
| Max. P | 0.680347 |

| Location | 797,274 – 797,372 |
|---|---|
| Length | 98 |
| Sequences | 10 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 78.26 |
| Shannon entropy | 0.47088 |
| G+C content | 0.43011 |
| Mean single sequence MFE | -18.07 |
| Consensus MFE | -10.46 |
| Energy contribution | -10.36 |
| Covariance contribution | -0.10 |
| Combinations/Pair | 1.22 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.680347 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 797274 98 + 23011544 CACAAUGCAACUCAUGAAAUCGACGAAUCGAC--AACUCGACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGUC--AGCCGCACCUCCACGG .....(((......((((...((((..((((.--...))))..))))..))))(((((((...(((((.....))))))))).)--))..)))......... ( -19.00, z-score = -0.99, R) >dp4.chr4_group3 2755839 83 - 11692001 ----------------AA-UCGACAACUCGAC--AACUCGACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGUCCUGGAUGCACUCUCAUAG ----------------..-((((....)))).--.....((.((((..(((((..(((((...(((((.....))))))))))..)))))))))))...... ( -17.30, z-score = -1.98, R) >droAna3.scaffold_12916 1760623 96 + 16180835 CACAAUGCAACUCAUGCA-CCGCCGAAUCGAC--AACUCGACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGUC--AGUGCCUCUCCUGUG- ((((.((((.....))))-..((.((((.(((--(........)))).))))((((((((...(((((.....))))))))).)--)))))......))))- ( -20.70, z-score = -1.72, R) >droEre2.scaffold_4929 851060 98 + 26641161 CACAAUGCAACUCAUGAAAUCGACGAAUCGAC--AACUCGACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGUC--AGCCACACCUCCACGG ...((((((.....((((...((((..((((.--...))))..))))..)))).))))))....(((((((((.....))))))--)))............. ( -18.60, z-score = -1.19, R) >droYak2.chr2L 779911 98 + 22324452 CACAAUGCAACUCAUGAAAUCGACGAAUCGAC--AACUCGACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGUC--AGCCGCACCUCCACGG .....(((......((((...((((..((((.--...))))..))))..))))(((((((...(((((.....))))))))).)--))..)))......... ( -19.00, z-score = -0.99, R) >droSec1.super_14 767255 98 + 2068291 CACAAUGCAACUCAUGAAAUCGACGAAUCGAC--AACUCGACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGUC--AGCCGCACCUCCACGG .....(((......((((...((((..((((.--...))))..))))..))))(((((((...(((((.....))))))))).)--))..)))......... ( -19.00, z-score = -0.99, R) >droSim1.chr2L 798719 98 + 22036055 CACAAUGCAACUCAUGAAAUCGACGAAUCGAC--AACUCGACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGUC--AGCCGCACCUCCACGG .....(((......((((...((((..((((.--...))))..))))..))))(((((((...(((((.....))))))))).)--))..)))......... ( -19.00, z-score = -0.99, R) >droVir3.scaffold_12963 19319716 97 + 20206255 ---CACAAUGCUGAACUCUUGAACGAAUUGAACAAACUCAACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGCC--AACACCACAUUCACAU ---....(((.((((....((((((..((((......))))..))))))......(((((...(((((.....)))))))))).--.........))))))) ( -16.70, z-score = -1.91, R) >droMoj3.scaffold_6500 8463842 97 - 32352404 ---CACUACGCUCAACUCUCAAACGAAUCGAACAAACUCGACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGCC--AUCGUCACAUUCACAG ---....(((..........(((((..((((......))))..))))).......(((((...(((((.....)))))))))).--..)))........... ( -15.10, z-score = -1.63, R) >droGri2.scaffold_15252 7195213 94 + 17193109 ---CAAUGCGCUCAGCACUU---CGAAUGAAACGAGCUCGAUGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGCC--AACACCAUAUUCACCA ---...(((.....)))...---.(((((((....((.....)).)))))))...(((((...(((((.....)))))))))).--................ ( -16.30, z-score = -0.45, R) >consensus CACAAUGCAACUCAUGAAAUCGACGAAUCGAC__AACUCGACGUGUUUAUUCACUGCAUUUCCUGUUGAUAUUCAACAAAUGUC__AGCCGCACCUCCACGG .....................((((..((((......))))..))))........(((((...(((((.....))))))))))................... (-10.46 = -10.36 + -0.10)


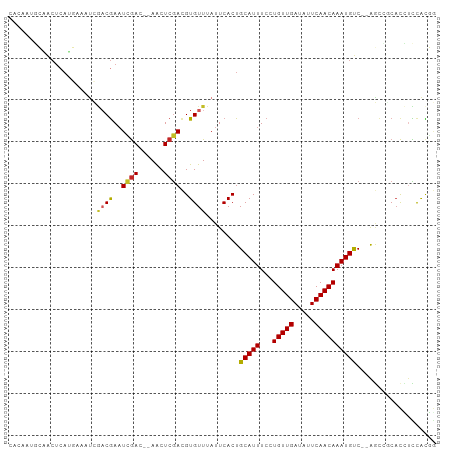
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:06:54 2011