
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,774,387 – 20,774,495 |
| Length | 108 |
| Max. P | 0.551186 |

| Location | 20,774,387 – 20,774,495 |
|---|---|
| Length | 108 |
| Sequences | 13 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 59.85 |
| Shannon entropy | 0.92804 |
| G+C content | 0.47304 |
| Mean single sequence MFE | -26.98 |
| Consensus MFE | -9.06 |
| Energy contribution | -9.29 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.50 |
| Mean z-score | -0.69 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.551186 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 20774387 108 - 27905053 CGAUCCAAACUUAUAGGCGGCUAUGAUGGAGCCAAAGAUGGCCACGCUGCAAGUGACUAAUCGAGAGGAAUGAAAUU---AAUAUCCAUCUGUAAUCGGAUU--GGAGGAGCA- ...((((...((((((....))))))))))((((....))))...(((.(..........((....)).........---....((((((((....)))).)--)))).))).- ( -25.30, z-score = -0.06, R) >droSim1.chr3R 20606288 108 - 27517382 CGAUCCAAAUUUGUAGGCGGCUAUGAUGGAGCCAAAGAUGGCCACGCUGCAGGUGACUGAUCAAGAGGAAUGAAAUU---AUUAUCCAUCUGCAAUCGGUUU--GGAGGAGCA- ...((((.(((((((((.((((((..((....))...)))))).).))))))))(((((((..((((((.(((....---.)))))).)))...))))))))--)))......- ( -32.00, z-score = -1.04, R) >droSec1.super_0 21116723 108 - 21120651 CGAUCCAAAUUUGUAGGCGGCUAUGAUGGAGCCAAAGAUGGCCACGCUGCAGGUGACUGAUCAGGAGGAAUGAAAUU---AUUAUCCAUCUGUAAUCGGUUU--GGAGGAGCA- ...((((.(((((((((.((((((..((....))...)))))).).))))))))((((((((((..(((.(((....---.))))))..)))..))))))))--)))......- ( -33.20, z-score = -1.52, R) >droYak2.chr3R 3106595 108 + 28832112 CGAUCCAAAUUUGUAGGCGGCUAUGAUGGAACCGAAGAUGGCCACGCUGCAAGUAACUGUUUAAAAGGAAUGCAAUU---AUUAUCCAUCCGUAAUCGGGAU--AGUGGAGCA- .(.(((..(((((((((.((((((..((....))...)))))).).)))))))).((((((.....(((........---....)))..(((....))))))--)))))).).- ( -28.70, z-score = -0.87, R) >droEre2.scaffold_4820 3190386 108 + 10470090 CGAUCCAAAUUUGUAGGCGGCUAUGAUGGAGCCGAAGAUAGCUACGCUGCAUGUAACUGAUCGGAAGGAAUGAAAUU---AUUAUCCAUCUGUAGUCGGUCU--AGUGCAGCG- ..........((((...(((((.......)))))...))))...((((((((...((((((((((.(((.(((....---.)))))).))))..))))))..--.))))))))- ( -34.70, z-score = -2.23, R) >droAna3.scaffold_13340 4817298 114 + 23697760 GGACCCAAAUUUAUAGGCGGCGAUGACAGAUCCGAUAAUGGCCACGCUGCAGGUAACUAAGGAUAGCAAAAGGAUUAGCAAUCAUUAUCCUAUACCCUCAGAUAAAUGAAAGCC ((......((((((..((((((.((.((..........))..)))))))).((((....(((((((......(((.....))).))))))).)))).....)))))).....)) ( -21.80, z-score = 0.41, R) >dp4.chr2 19832371 103 + 30794189 CGACCCAAACUUGUAGGCGGCAAUGAUGGACCCAAAGAUGGCCACGCUGCAGGUAACUAAGA-------GUGGGUAU-CCAACCUCCAUUAGUGACCAUUCG--ACUGUAGUG- (((.((.........)).((((.(((((((......((((.((((.((..((....)).)).-------)))).)))-).....))))))).)).))..)))--.........- ( -25.40, z-score = 0.49, R) >droPer1.super_0 9505769 103 - 11822988 GAACCCAAACUUGUAGGCGGCAAUUAUGGACCCAAAGAUGGCCACGCUGCAGGUAACUAAGA-------GUGGGUAU-CCAACCUCCAUUAGUGACCAUUCG--ACUGUAGUG- ........(((((((((.(((.(((.((....))..))).))).).)))))))).((((.((-------((((....-.................)))))).--....)))).- ( -25.00, z-score = 0.31, R) >droWil1.scaffold_181130 4011464 109 + 16660200 UGAGCCAAAUUUGUAAGCCGCAAUAAUGGAACCAAAGAUGGCCACGCCACAGGUAACUACACAUGUGAUAUGAAAAUUUAAGCUCAAAUUUAUAACUAAAAA--ACUUUUC--- (((((.((((((......((((.....(..(((...(.((((...))))).)))..)......))))......))))))..)))))................--.......--- ( -16.70, z-score = -0.21, R) >droVir3.scaffold_13047 12891208 102 - 19223366 GGCACCGAACUUGUAGGCGGCUAUGAUGGCACCAACAAUGGCCACGCUGCAAGUGACUAAACA---G--AUGGAGAU--CAACGUGCUGCGGCAGCCGCAUUA-ACAAGU---- (((((...(((((((((.((((((..((....))...)))))).).)))))))).........---(--((....))--)...)))))((((...))))....-......---- ( -33.10, z-score = -0.92, R) >droMoj3.scaffold_6540 5184860 108 + 34148556 GGAGCCGAACUUGUAGGCAGCUAUGCCGGCACCCACAAUGGCCACGCUGCAAGUGACUAAACA---GAAAUAGAUAU--ACUCGUCUUUCGACAUCCACAGUACAUAAGAGCA- (((..((((((((((((((....))))(((..........)))....))))))).........---.....((((..--....)))).)))...)))................- ( -23.70, z-score = 0.11, R) >droGri2.scaffold_15074 5938937 103 + 7742996 GGAACCAAACUUGUAGGCGGCGAUAACCGCACCAAUGAUGGCCACACUGCAAGUAACUAAACC---G--GUUCAAAU--AAACAACAAAUAAUAAACACAA---AUAAUACCA- .(((((..((((((((((((......)))).(((....))).....)))))))).........---)--))))....--......................---.........- ( -21.80, z-score = -2.60, R) >anoGam1.chr2R 41929664 84 + 62725911 CAUCCCAUAACGGUACGCGGUCAGCAGCGUGGUGACCGCGGACAGGCUGCAGGUCG-UAACGAUGCCAAACAGUACCGCCUUCCC----------------------------- ..........((((((...((..(((.(((..(((((((((.....)))).)))))-..))).)))...)).)))))).......----------------------------- ( -29.40, z-score = -0.86, R) >consensus CGAUCCAAACUUGUAGGCGGCUAUGAUGGAACCAAAGAUGGCCACGCUGCAGGUAACUAAUCA___GGAAUGGAAAU___AUCAUCCAUCUGUAACCGGAUU__AGAGGAGCA_ ........(((((((((.(((...................))).).))))))))............................................................ ( -9.06 = -9.29 + 0.24)


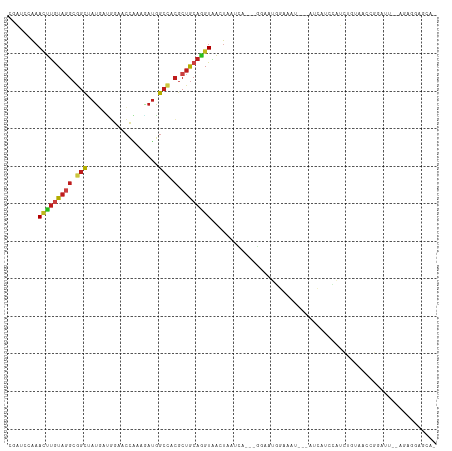
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:41:50 2011