
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,725,770 – 20,725,880 |
| Length | 110 |
| Max. P | 0.833117 |

| Location | 20,725,770 – 20,725,880 |
|---|---|
| Length | 110 |
| Sequences | 7 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 74.84 |
| Shannon entropy | 0.49345 |
| G+C content | 0.50519 |
| Mean single sequence MFE | -27.55 |
| Consensus MFE | -13.57 |
| Energy contribution | -15.72 |
| Covariance contribution | 2.15 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.833117 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
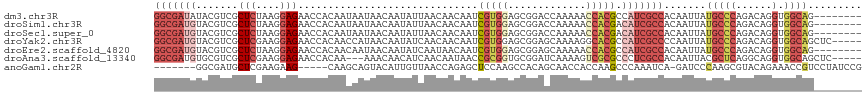
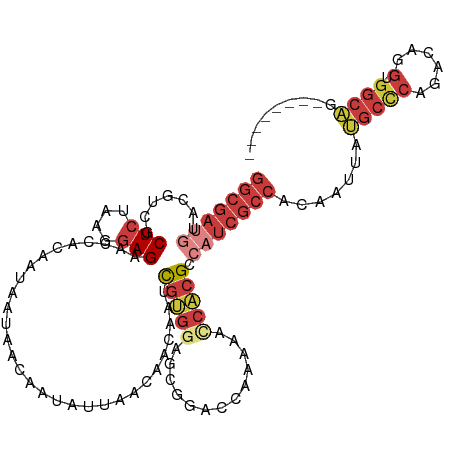
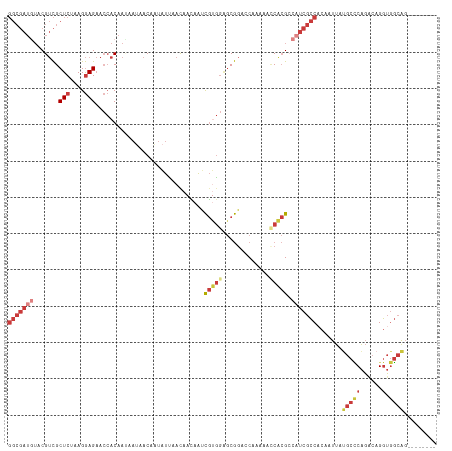
>dm3.chr3R 20725770 110 - 27905053 GGCGAUAUACGUCGCUCUAAGGAGAACCACAAUAAUAACAAUAUUAACAACAAUCGUGGAGCGGACCAAAAACCACGCCAUCGCCACAAUUAUGCCCAGACAGGUGGCAG-------- ((((.........((((((.((....))....(((((....)))))..........))))))((........)).))))...(((((.....((......)).)))))..-------- ( -26.40, z-score = -2.23, R) >droSim1.chr3R 20560549 110 - 27517382 GGCGAUGUACGUCGCUCUAAGGAGAACCACAAUAAUAACAAUAUUAACAACAAUCGUGGAGCGGACCAAAAACCACGACAUCGCCACAAUUAUGCCCAGACAGGUGGCAG-------- ((((((((.(((.((((((.((....))....(((((....)))))..........))))))((........))))))))))))).......(((((......).)))).-------- ( -32.30, z-score = -3.99, R) >droSec1.super_0 21073640 110 - 21120651 GGCGAUGUACGUCGCUCUAAGGAGAACCACAAUAAUAACAAUAUUAACAACAAUCGUGGAGCGGACCAAAAACCACGACAUCGCCACAAUUAUGCCCAGACAGGUGGCAG-------- ((((((((.(((.((((((.((....))....(((((....)))))..........))))))((........))))))))))))).......(((((......).)))).-------- ( -32.30, z-score = -3.99, R) >droYak2.chr3R 3062225 113 + 28832112 GGCGAUGUACGUCGCUCGAAGGAGAACCACAACCAUAACAAUAUCAACAACAAUCGUGGAGCGGAGCAAAAGGCACGCCAUCGCCCCAAUUAUGCCCAGACAGGUGGCAGCUC----- (((((((.......(((....)))..((((.........................)))).(((..((.....)).)))))))))).......(((((......).))))....----- ( -29.01, z-score = -0.48, R) >droEre2.scaffold_4820 3146291 110 + 10470090 GGCGAUGUACGUCGCUCUAAGGAGAACCACAACAAUAACAAUAUCAAUAACAAUCGUGGAGCGGAGCAAAAACCACGCCAUCGCCACAAUUAUGCCCAGACAGGUGGCAG-------- (((((((..(((.((((((..((..............................)).))))))((........))))).))))))).......(((((......).)))).-------- ( -26.71, z-score = -1.47, R) >droAna3.scaffold_13340 4773593 110 + 23697760 GGCGAUGUGCGUCGCUCGAAGGAGAACCACAA---AAACAACAUCAACAAUAACCGCGGUGCGGAUCAAAAGUCGCGCCCUCGCCACAAUUACGCUCAGGCAGGUGGCAGCUC----- (((((((((.((..(((....))).)))))).---......................(((((((........))))))).))(((((......((....))..))))).))).----- ( -31.40, z-score = -0.13, R) >anoGam1.chr2R 48074682 105 + 62725911 -------GGCGAUGCUCGAAGAAG-----CAAGCAGUACAUUGUUAACCAGAGCUCCAAGCCACAGCAACCACCAAGCCCAAAUCA-GAUCCCAAGCGUACAGAAACCGUCCUAUCCG -------(((((.((((.....((-----(((........))))).....))))))...((....)).........))).......-............................... ( -14.70, z-score = 0.02, R) >consensus GGCGAUGUACGUCGCUCUAAGGAGAACCACAAUAAUAACAAUAUUAACAACAAUCGUGGAGCGGACCAAAAACCACGCCAUCGCCACAAUUAUGCCCAGACAGGUGGCAG________ (((((((.......(((....)))..............................(((((.............))))).))))))).......(((((......).))))......... (-13.57 = -15.72 + 2.15)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:41:44 2011