
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,435,956 – 20,436,038 |
| Length | 82 |
| Max. P | 0.879830 |

| Location | 20,435,956 – 20,436,038 |
|---|---|
| Length | 82 |
| Sequences | 4 |
| Columns | 82 |
| Reading direction | reverse |
| Mean pairwise identity | 80.20 |
| Shannon entropy | 0.32511 |
| G+C content | 0.56799 |
| Mean single sequence MFE | -30.27 |
| Consensus MFE | -15.70 |
| Energy contribution | -19.20 |
| Covariance contribution | 3.50 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.58 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.879830 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
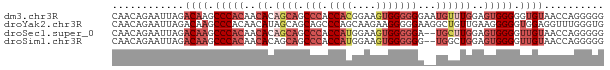
>dm3.chr3R 20435956 82 - 27905053 CAACAGAAUUAGACAAGCCCACAACACAGCAGCCCACCACGGAAGUGGGGGGAAUGUUUGGAGUGGGGGUGUAACCAGGGGG ............(((..(((((..((.((((.(((.((((....)))))))...))))))..)))))..))).......... ( -31.00, z-score = -2.71, R) >droYak2.chr3R 2768874 82 + 28832112 CAACAGAAUUAGACAAGCCCACAACAUAGCAGCAGCCCAGCAAGAAGGGGGAAGGCUGUUGAAGGGGGUGGAGGUUUGGGUG .............(((((((((..(.(((((((..(((..(.....).)))...)))))))...)..)))..)))))).... ( -25.30, z-score = -2.13, R) >droSec1.super_0 20769594 80 - 21120651 CAACAGAAUUAGACAAGCCCACAACACAGCAGCCCACCAUGGAAGUGGGGGA--UGCUUGGAGUGGGGUUGUAACCAGGGGG ............((((.(((((..((.((((.(((.((((....))))))).--))))))..))))).)))).......... ( -31.80, z-score = -2.86, R) >droSim1.chr3R 20273708 80 - 27517382 CAACAGAAUUAGACAAGCCCACAACACAGCAGCCCACCAUGGAAGUGGGGGG--UGGCUGGAGUGGGGUUGUAACCAGGGGG ............((((.(((((....((((.((((.((((....)))).)))--).))))..))))).)))).......... ( -33.00, z-score = -2.62, R) >consensus CAACAGAAUUAGACAAGCCCACAACACAGCAGCCCACCAUGGAAGUGGGGGA__UGUUUGGAGUGGGGUUGUAACCAGGGGG ............((((.(((((..((.((((.(((.((((....)))))))...))))))..))))).)))).......... (-15.70 = -19.20 + 3.50)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:41:12 2011