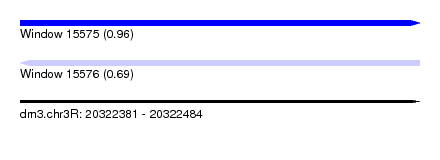

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,322,381 – 20,322,484 |
| Length | 103 |
| Max. P | 0.962966 |

| Location | 20,322,381 – 20,322,484 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 78.11 |
| Shannon entropy | 0.29177 |
| G+C content | 0.57018 |
| Mean single sequence MFE | -35.70 |
| Consensus MFE | -24.30 |
| Energy contribution | -25.63 |
| Covariance contribution | 1.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.53 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.71 |
| SVM RNA-class probability | 0.962966 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
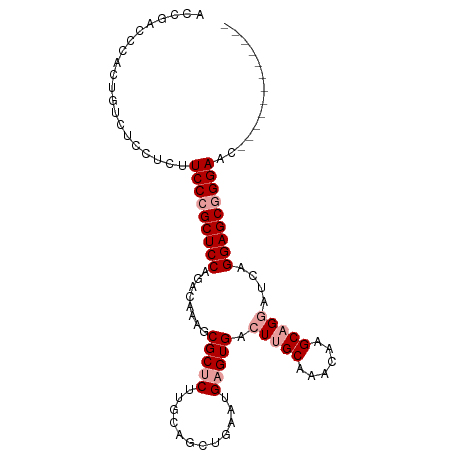
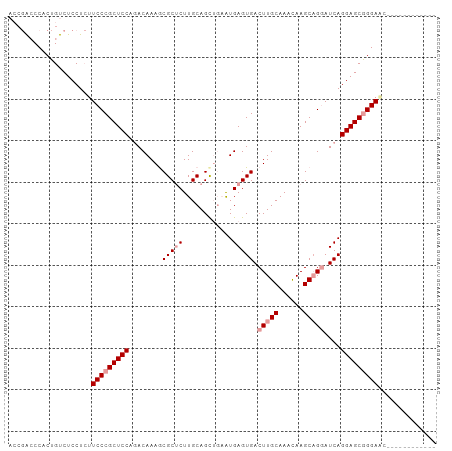
>dm3.chr3R 20322381 103 + 27905053 ACCGACCCACUGUCUCCUCUUCCCGCUCCACACAAAACGCUCUUGCAGCUGAAUGAGUGACUUGCAAACAAGCAGUAUCAGGAGCAGGAUCAGGAGCGGGAGC ...(((.....))).....((((((((((.........(((((((..((((..((..((.....))..))..))))..))))))).......)))))))))). ( -35.79, z-score = -2.02, R) >droSec1.super_0 20658171 91 + 21120651 ACCGACCCACUGUCUCCUCUUCCCGCUCCAGACAAAGCGCUCUUGCAGCGCUGUGAGUGACUUGCAAACAAGCAGGAUCAGGAGCGGGAAC------------ ...(((.....))).....((((((((((...((.((((((.....)))))).))..((((((((......))))).))))))))))))).------------ ( -39.80, z-score = -3.93, R) >droEre2.scaffold_4820 2733140 87 - 10470090 ----GGCCACUGUCUCCUGUUCCCGCUCCAGACAAAGCGCCCUUGCAGUUGAAUGAGUGACUUGCAAAUAAGCCGGAUCAGGAGCGGGAGC------------ ----(((....)))....(((((((((((.(((...((....((((((((........))).)))))....))..).)).)))))))))))------------ ( -31.50, z-score = -1.63, R) >consensus ACCGACCCACUGUCUCCUCUUCCCGCUCCAGACAAAGCGCUCUUGCAGCUGAAUGAGUGACUUGCAAACAAGCAGGAUCAGGAGCGGGAAC____________ ....................(((((((((........(((((............))))).(((((......)))))....))))))))).............. (-24.30 = -25.63 + 1.33)

| Location | 20,322,381 – 20,322,484 |
|---|---|
| Length | 103 |
| Sequences | 3 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 78.11 |
| Shannon entropy | 0.29177 |
| G+C content | 0.57018 |
| Mean single sequence MFE | -36.53 |
| Consensus MFE | -29.26 |
| Energy contribution | -28.93 |
| Covariance contribution | -0.33 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.689302 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
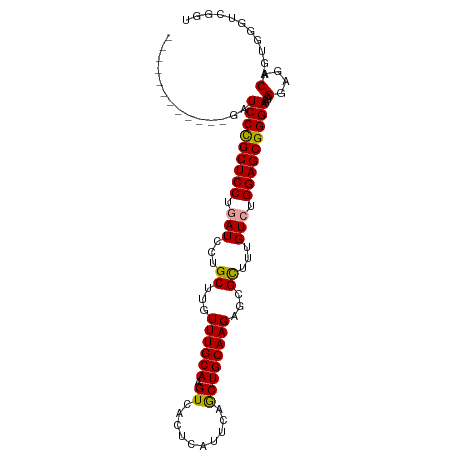
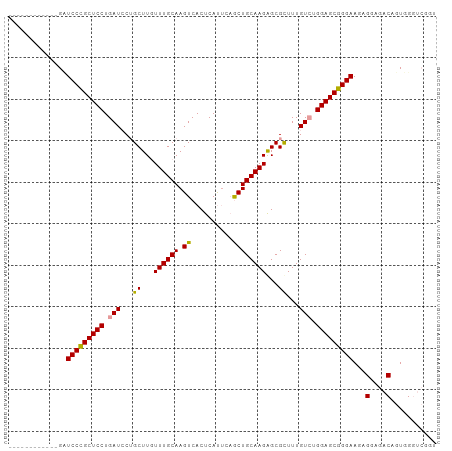
>dm3.chr3R 20322381 103 - 27905053 GCUCCCGCUCCUGAUCCUGCUCCUGAUACUGCUUGUUUGCAAGUCACUCAUUCAGCUGCAAGAGCGUUUUGUGUGGAGCGGGAAGAGGAGACAGUGGGUCGGU ((((((((((((..(((((((((.......(((((....))))).((.((....(((.....)))....)).)))))))))))..))))....)))))..))) ( -38.40, z-score = -0.94, R) >droSec1.super_0 20658171 91 - 21120651 ------------GUUCCCGCUCCUGAUCCUGCUUGUUUGCAAGUCACUCACAGCGCUGCAAGAGCGCUUUGUCUGGAGCGGGAAGAGGAGACAGUGGGUCGGU ------------.((((((((((.((....(((((....)))))....((.((((((.....)))))).)))).)))))))))).....(((.....)))... ( -37.60, z-score = -1.41, R) >droEre2.scaffold_4820 2733140 87 + 10470090 ------------GCUCCCGCUCCUGAUCCGGCUUAUUUGCAAGUCACUCAUUCAACUGCAAGGGCGCUUUGUCUGGAGCGGGAACAGGAGACAGUGGCC---- ------------(((((((((((.(((..(((...((((((.((..........))))))))...)))..))).)))))))))...(....)....)).---- ( -33.60, z-score = -1.75, R) >consensus ____________GAUCCCGCUCCUGAUCCUGCUUGUUUGCAAGUCACUCAUUCAGCUGCAAGAGCGCUUUGUCUGGAGCGGGAAGAGGAGACAGUGGGUCGGU ..............(((((((((.(((...((...((((((.((..........))))))))...))...))).)))))))))...(....)........... (-29.26 = -28.93 + -0.33)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:40:52 2011