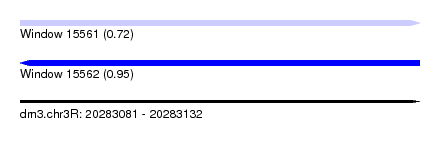

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,283,081 – 20,283,132 |
| Length | 51 |
| Max. P | 0.953336 |

| Location | 20,283,081 – 20,283,132 |
|---|---|
| Length | 51 |
| Sequences | 4 |
| Columns | 52 |
| Reading direction | forward |
| Mean pairwise identity | 90.29 |
| Shannon entropy | 0.15602 |
| G+C content | 0.53658 |
| Mean single sequence MFE | -17.40 |
| Consensus MFE | -14.57 |
| Energy contribution | -14.82 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.724856 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
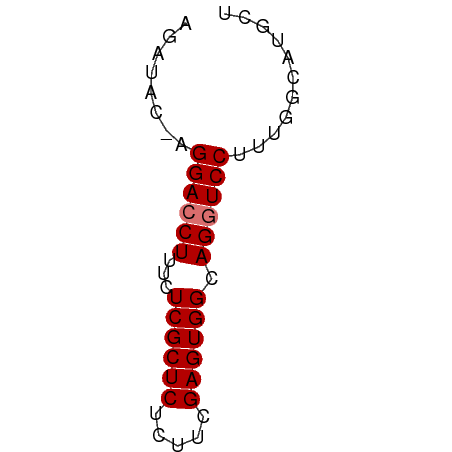
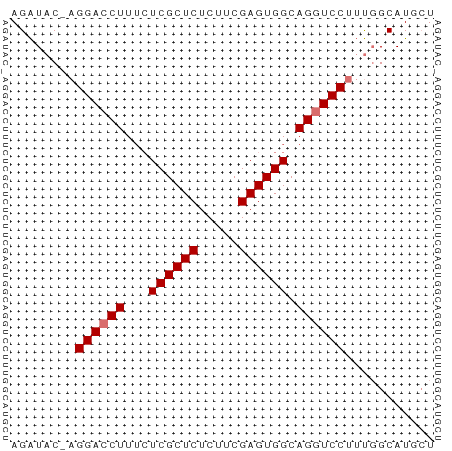
>dm3.chr3R 20283081 51 + 27905053 AGAUAC-AGGACCUUUCUCGCUCUCUUCGAGUGGCAGGUCCUUUGGCAUGAU .(.((.-(((((((...((((((.....)))))).))))))).)).)..... ( -17.10, z-score = -1.73, R) >droSec1.super_0 20619669 52 + 21120651 AGAUACCAGGACCUUUCUCGCUCUCUUCGAGUGGCAGAUCCCUUGGCAUGCU .....(((((...(((.((((((.....)))))).)))...)))))...... ( -12.20, z-score = 0.20, R) >droYak2.chr3R 2612094 51 - 28832112 AGAUAC-AGGACCUUGCUCGCUCUCUUCGAGUGGCAGGUCCUUUCGCAUGCU .((...-(((((((.((.(((((.....)))))))))))))).))....... ( -21.70, z-score = -3.25, R) >droEre2.scaffold_4820 2693859 51 - 10470090 AGGUAC-AGGACCUUUAUCGCUCUCUUCGAGUGGCAGGUCCUACGGCAUGCU ..((..-(((((((...((((((.....)))))).)))))))...))..... ( -18.60, z-score = -2.20, R) >consensus AGAUAC_AGGACCUUUCUCGCUCUCUUCGAGUGGCAGGUCCUUUGGCAUGCU ........((((((...((((((.....)))))).))))))........... (-14.57 = -14.82 + 0.25)

| Location | 20,283,081 – 20,283,132 |
|---|---|
| Length | 51 |
| Sequences | 4 |
| Columns | 52 |
| Reading direction | reverse |
| Mean pairwise identity | 90.29 |
| Shannon entropy | 0.15602 |
| G+C content | 0.53658 |
| Mean single sequence MFE | -17.00 |
| Consensus MFE | -15.30 |
| Energy contribution | -14.92 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.09 |
| Structure conservation index | 0.90 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.59 |
| SVM RNA-class probability | 0.953336 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
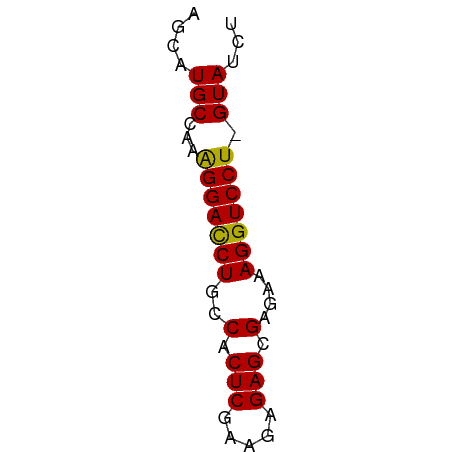
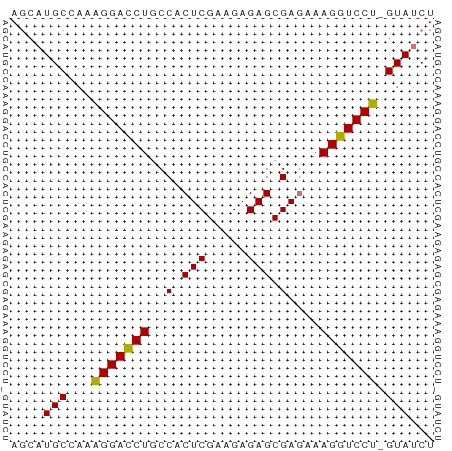
>dm3.chr3R 20283081 51 - 27905053 AUCAUGCCAAAGGACCUGCCACUCGAAGAGAGCGAGAAAGGUCCU-GUAUCU ...((((...(((((((....((((.......))))..)))))))-)))).. ( -15.90, z-score = -2.22, R) >droSec1.super_0 20619669 52 - 21120651 AGCAUGCCAAGGGAUCUGCCACUCGAAGAGAGCGAGAAAGGUCCUGGUAUCU ((.((((((..((((((....((((.......))))..)))))))))))))) ( -17.30, z-score = -1.44, R) >droYak2.chr3R 2612094 51 + 28832112 AGCAUGCGAAAGGACCUGCCACUCGAAGAGAGCGAGCAAGGUCCU-GUAUCU ((.((((...((((((((((.(((.....))).).)).)))))))-)))))) ( -19.20, z-score = -2.66, R) >droEre2.scaffold_4820 2693859 51 + 10470090 AGCAUGCCGUAGGACCUGCCACUCGAAGAGAGCGAUAAAGGUCCU-GUACCU .........((((((((..(.(((.....))).)....)))))))-)..... ( -15.60, z-score = -2.02, R) >consensus AGCAUGCCAAAGGACCUGCCACUCGAAGAGAGCGAGAAAGGUCCU_GUAUCU ....(((...(((((((..(.(((.....))).)....))))))).)))... (-15.30 = -14.92 + -0.38)
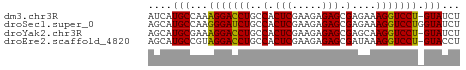

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:40:41 2011