
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,262,985 – 20,263,067 |
| Length | 82 |
| Max. P | 0.807671 |

| Location | 20,262,985 – 20,263,067 |
|---|---|
| Length | 82 |
| Sequences | 12 |
| Columns | 88 |
| Reading direction | reverse |
| Mean pairwise identity | 70.97 |
| Shannon entropy | 0.61691 |
| G+C content | 0.36352 |
| Mean single sequence MFE | -12.68 |
| Consensus MFE | -8.03 |
| Energy contribution | -8.17 |
| Covariance contribution | 0.13 |
| Combinations/Pair | 1.50 |
| Mean z-score | -0.57 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.807671 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3R 20262985 82 - 27905053 UUUUCCGUGCAAUCCAACUUAAUGAUGCGCGAACUAUUAAGCGCAUUAAGUGCGAUUUUUG----CCUUUAUUUCCUU-UUGUCCAC- ......(..(((..........((((((((..........))))))))((.((((...)))----)))..........-)))..)..- ( -15.00, z-score = -1.51, R) >droSim1.chr3R 20103540 82 - 27517382 UUUUCCGUGCAAUCCAACUUAAUGAUGCGCGAACUAUUAAGCGCAUUAAGUGCGAUUUUUG----CCUUUAUUUCCUU-UUGUCCAC- ......(..(((..........((((((((..........))))))))((.((((...)))----)))..........-)))..)..- ( -15.00, z-score = -1.51, R) >droSec1.super_0 20599807 82 - 21120651 UUUUCCGUGCAAUCCAACUUAAUGAUGCGCGAACUAUUAAGUGCAUUAAGUGCGAUUUUUG----CCUUUAUUUCCUU-UUGUCCAC- ......(..(((..........((((((((..........))))))))((.((((...)))----)))..........-)))..)..- ( -13.10, z-score = -0.99, R) >droYak2.chr3R 2591285 84 + 28832112 UUUUCCCUGCAAUCCAACUUAAUGAUGCGCGAACUAUUAAGCGCAUUAAGAGCGAUUUUUG---CCCUUUAUUUCCUU-UUUGCCAAC ........((((..........((((((((..........))))))))((.((((...)))---).))..........-.)))).... ( -15.50, z-score = -2.25, R) >droEre2.scaffold_4820 2673860 82 + 10470090 UUUUCCCUGCAAUCCAACUUAAUGAUGCGCGAACUAUUAAACGCAUUAAGCGCGAUUUUUG----CCUUUAUUUCCUU-UUGUCGAC- ......(.((((..........(((((((............)))))))((.((((...)))----)))..........-)))).)..- ( -9.20, z-score = 0.29, R) >droAna3.scaffold_13088 303281 84 - 569066 UUUUUUGUGCAAUCCAACUUAAUGAUGCACGAACUAUUAAGCGUAUUAAAAGCGAUUUUUG---CCUUUUACCUUUUUGCUGUCGAC- ...((((((((.((.........))))))))))......((((...(((((((((...)))---.))))))......))))......- ( -12.70, z-score = -0.64, R) >dp4.chr2 12254002 86 + 30794189 CUUUUCAUGCAAUUCAACUUAAUGAUGCGCGAACUAUUAAGCGCAUUAAGAGCCAUUUUUUGUUGCCUUUACCUCCUU-UUGUCUGC- .....((.((((..........((((((((..........))))))))((((.((........)).))))........-)))).)).- ( -14.80, z-score = -0.92, R) >droPer1.super_3 6104065 86 + 7375914 CUUUUCAUGCAAUUCAACUUAAUGAUGCGCGAACUAUUAAGCGCAUUAAGAGCCAUUUUUUGUUGCCCUUACCUCCUU-UUGUCUGC- .....((.((((...........(((((((..........)))))))(((.((.((.....)).)).)))........-)))).)).- ( -13.90, z-score = -0.70, R) >droWil1.scaffold_181108 270020 66 + 4707319 UUUUUUGUGAAACUCGAUUUAAUGAUGCGCAAACUAUUAAGCUCUUUGAAAACAUUUUUU-----GUUCCA----------------- ...((((((....(((......)))..)))))).................((((.....)-----)))...----------------- ( -4.20, z-score = 1.43, R) >droVir3.scaffold_13047 3622014 75 - 19223366 CUCUUUAUGCAAUUGGAUUUAAUGAUGCGCGAACUAUUAAGCGCAUUAAAAGC-AUUAAAAUCGUAUUUUGUCCGC------------ ........((((.(((.(((((((((((((..........))))))......)-)))))).)))....))))....------------ ( -13.60, z-score = -0.16, R) >droMoj3.scaffold_6540 20629977 75 - 34148556 CUCUUUAUGAAAUUGGAUUUAAUGAUGCGCGAACUAUUAAGCACAUUAAAAGC-AUUAAAAGCAUUUUUUGCCAGC------------ ............((((.(((((((.(((............)))))))))).((-.......))........)))).------------ ( -11.10, z-score = 0.39, R) >droGri2.scaffold_14906 277435 79 - 14172833 CUCCUUAUGCAAUUGGAUUUAAUGAUGUGCGAACUAUUAAGCGCAUUAAGAGC-AUUAAAAGUCCUUUUUAUUUUUGCGC-------- ........((((..((((((((((((((((..........))))))......)-)))))..)))).........))))..-------- ( -14.10, z-score = -0.21, R) >consensus UUUUUCAUGCAAUCCAACUUAAUGAUGCGCGAACUAUUAAGCGCAUUAAGAGCGAUUUUUG____CCUUUAUUUCCUU_UUGUC_AC_ ......................((((((((..........))))))))........................................ ( -8.03 = -8.17 + 0.13)

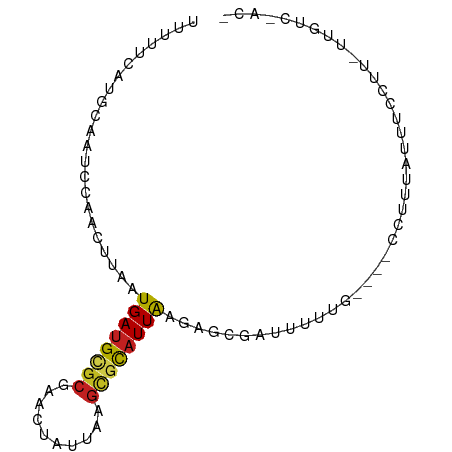

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:40:34 2011