
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,248,446 – 20,248,537 |
| Length | 91 |
| Max. P | 0.840217 |

| Location | 20,248,446 – 20,248,537 |
|---|---|
| Length | 91 |
| Sequences | 8 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 75.80 |
| Shannon entropy | 0.43697 |
| G+C content | 0.42150 |
| Mean single sequence MFE | -21.38 |
| Consensus MFE | -12.40 |
| Energy contribution | -12.06 |
| Covariance contribution | -0.34 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.87 |
| SVM RNA-class probability | 0.840217 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
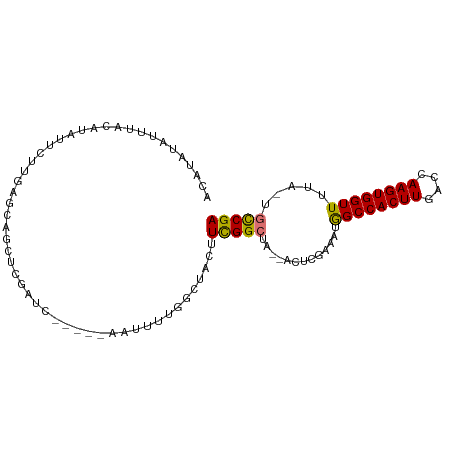
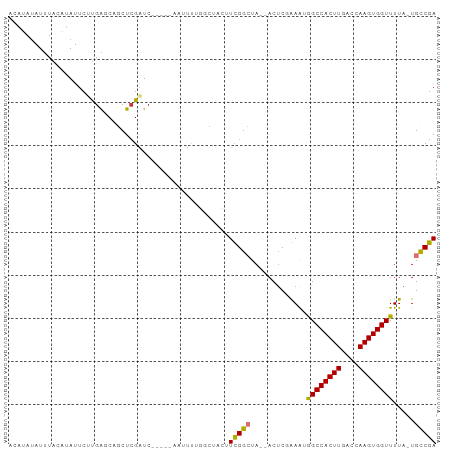
>dm3.chr3R 20248446 91 + 27905053 ACAUAUAUUUACAUAUUCUUGAGCAGCUCGAUC-----AAUUCUGGCUACUUCGGCUA--ACUCGGAAUGGCCACUUGACCAAGUGGUUUUA-UGCCGA .................((.(((.((((.((..-----...)).)))).))).))...--..((((...((((((((....))))))))...-..)))) ( -22.30, z-score = -1.63, R) >droPer1.super_3 6089064 91 - 7375914 ----AUAUUUA---AUUCAUGACCAACUCGUUCAAUUUUCCUUUCCUUCUGUUGGACAAAACUC-AAGCAGCCACUUGCCAAAGUGGUUUUAAUGCCGA ----.......---...............(((((((..............))))))).......-..((((((((((....))))))).....)))... ( -15.74, z-score = -1.09, R) >dp4.chr2 12239023 91 - 30794189 ----AUAUUUA---AUUCAUGACCAACUCGUUCAAUUUUCCUUUCCUUCUGUUGGACAAAACUC-AAGCAGCCACUUGCCAAAGUGGUUUUAAUGCCGA ----.......---...............(((((((..............))))))).......-..((((((((((....))))))).....)))... ( -15.74, z-score = -1.09, R) >droAna3.scaffold_13088 288296 83 + 569066 ----ACAUAUA---AUUCAUGGCCAGCUGGAUG-----UGCUUCACUCCUCUUGGCCA--ACUC-AAACAGCCACUUGGCCAAGUGGUUUUA-UGCCGA ----.((((..---.....(((((((..(((.(-----((...))))))..)))))))--....-....((((((((....)))))))).))-)).... ( -27.40, z-score = -2.50, R) >droSec1.super_0 20585292 91 + 21120651 ACAUAUAUUUACAUAUUCUUGAGCAGCUCGAUC-----AAUUCUGGCUACUUCGGCUA--ACACGGAAUGGCCACUUGACCAAGUGGUUUUA-UGCCGA .................((.(((.((((.((..-----...)).)))).))).))...--...(((...((((((((....))))))))...-..))). ( -21.90, z-score = -1.60, R) >droEre2.scaffold_4820 2658675 91 - 10470090 ACAUAUAUUUACAUAUUCCUGAGCAGCUCGAUC-----AAUUUUGGCUACUUCGGCUA--ACUCGGAAUGGCCACUUGACCAAGUGGUUUUA-UGCCGA .................((.(((.(((.(((..-----....)))))).))).))...--..((((...((((((((....))))))))...-..)))) ( -23.10, z-score = -1.86, R) >droYak2.chr3R 2576082 91 - 28832112 ACAUAUAUUUACAUAUUCUCGAGCAGCUCGAUC-----CAUUUUGGUUCCUUUGGCUA--ACUCGAAAUGGCCACUUGACCAAGUGGUUUUA-UGCCGA ..................(((.(((.......(-----(((((.((((....((((((--........))))))...)))))))))).....-)))))) ( -22.70, z-score = -2.05, R) >droSim1.chr3R 20085549 91 + 27517382 ACAUAUAUUUACAUAUUCUUGAGCAGCUCGAUC-----AAUUCUGGCUACUUCGGCUA--ACUCGGAAUGGCCACUUGACCAAGUGGUUUUA-UGUCGA (((((........((((((.(((.((((.((..-----...)).)))).(....)...--.)))))))))(((((((....)))))))..))-)))... ( -22.20, z-score = -1.70, R) >consensus ACAUAUAUUUACAUAUUCUUGAGCAGCUCGAUC_____AAUUUUGGCUACUUCGGCUA__ACUCGAAAUGGCCACUUGACCAAGUGGUUUUA_UGCCGA ...................................................(((((.............((((((((....)))))))).....))))) (-12.40 = -12.06 + -0.34)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:40:30 2011