
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,202,209 – 20,202,314 |
| Length | 105 |
| Max. P | 0.666610 |

| Location | 20,202,209 – 20,202,311 |
|---|---|
| Length | 102 |
| Sequences | 5 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 75.57 |
| Shannon entropy | 0.41709 |
| G+C content | 0.44863 |
| Mean single sequence MFE | -26.88 |
| Consensus MFE | -14.78 |
| Energy contribution | -16.42 |
| Covariance contribution | 1.64 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.66 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.666610 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 20202209 102 + 27905053 GAGUCAUUGCGUAUCGAAG-----UAAGUGUUUCAGAUAGUUGGAAUGGCCACAGCUUCAGUUGCGGUUUCGAUGCUCAGUAUUCAUUUUGUCCAUUCGUUUUAGUA--------- ((((.((((.((((((((.-----.....(((((((....))))))).(((.((((....)))).))))))))))).))))))))......................--------- ( -26.70, z-score = -2.32, R) >droSim1.chr3R 20037082 115 + 27517382 GAGGCAUUGCGAAUCGAAGCCACGUAAGUGUUCCAGAUAGUUGGAGUCGCCACAGCUUCAGUUGCGGUUUCGAUGUUCAGUAUUGAUUCUGCCCAUUCGUUUUA-UACAUACAUUU (.((((....(((((((.(((((....))).....((..((((((..(((.((.......)).)))..))))))..)).)).))))))))))))..........-........... ( -29.80, z-score = -1.45, R) >droSec1.super_0 20540952 107 + 21120651 GAAGUAUUGCGUAUCGAAGCCACGUAAGUGUUCCAGAUAGUUGGAGUCGCCACAGCUUCAGUUGCAGUUUCGAUGUUCAGUAUUGAUUCUGCCGAUUCGUUUUAGUA--------- ..(.((..(((.(((((((((((....)))((((((....))))))........)))))....((((..((((((.....))))))..)))).))).)))..)).).--------- ( -31.50, z-score = -3.18, R) >droYak2.chr3R 2529543 86 - 28832112 GAGGCAUUGCAUCUCGAAGCCACGUAAGCGUUCCAGGU------------------UCCAGUUGCGGUUUCGAUGUUCACUAUUGAUUCCAUCCAUUCAUUUUU------------ ..((....(((.((.(((.(((((....)))....)))------------------)).)).)))((..((((((.....))))))..))..))..........------------ ( -17.60, z-score = -0.60, R) >droEre2.scaffold_4820 2612473 103 - 10470090 GAGGCAUUGCGGAUCGAAGCCACGUAAGUGUUCCAGAUAGUUGCAGUGGCCACAGCUUCAGUUGCGGUUUCGGUGCUCAGUAUUGAUUCCAUCCAAUCUUUUG------------- ..(((((((((.(((.....(((....))).....)))...)))))).))).((((....)))).((..(((((((...)))))))..)).............------------- ( -28.80, z-score = -0.75, R) >consensus GAGGCAUUGCGUAUCGAAGCCACGUAAGUGUUCCAGAUAGUUGGAGUCGCCACAGCUUCAGUUGCGGUUUCGAUGUUCAGUAUUGAUUCUGUCCAUUCGUUUUA_UA_________ .....((((.(((((((((((..((((.((........(((((.........))))).)).))))))))))))))).))))................................... (-14.78 = -16.42 + 1.64)



| Location | 20,202,215 – 20,202,314 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 74.10 |
| Shannon entropy | 0.44222 |
| G+C content | 0.43742 |
| Mean single sequence MFE | -25.58 |
| Consensus MFE | -13.34 |
| Energy contribution | -14.78 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.621619 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 20202215 99 + 27905053 UUGCGUAUCGAAG-----UAAGUGUUUCAGAUAGUUGGAAUGGCCACAGCUUCAGUUGCGGUUUCGAUGCUCAGUAUUCAUUUUGUCCAUUCGUUUUAGUAGAA----------- (((.((((((((.-----.....(((((((....))))))).(((.((((....)))).))))))))))).)))..............................----------- ( -21.80, z-score = -0.82, R) >droSim1.chr3R 20037088 115 + 27517382 UUGCGAAUCGAAGCCACGUAAGUGUUCCAGAUAGUUGGAGUCGCCACAGCUUCAGUUGCGGUUUCGAUGUUCAGUAUUGAUUCUGCCCAUUCGUUUUAUACAUACAUUUUAUACA ..((((((.((((((((....)))((((((....))))))........)))))....((((..((((((.....))))))..))))..))))))..................... ( -29.10, z-score = -2.04, R) >droSec1.super_0 20540958 104 + 21120651 UUGCGUAUCGAAGCCACGUAAGUGUUCCAGAUAGUUGGAGUCGCCACAGCUUCAGUUGCAGUUUCGAUGUUCAGUAUUGAUUCUGCCGAUUCGUUUUAGUAGCA----------- ..(((.(((((((((((....)))((((((....))))))........)))))....((((..((((((.....))))))..)))).))).)))..........----------- ( -29.90, z-score = -2.62, R) >droYak2.chr3R 2529549 83 - 28832112 UUGCAUCUCGAAGCCACGUAAGCGUUCCAGGU------------------UCCAGUUGCGGUUUCGAUGUUCACUAUUGAUUCCAUCCAUUCAUUUUUGAA-------------- ..(((.((.(((.(((((....)))....)))------------------)).)).)))((..((((((.....))))))..)).....((((....))))-------------- ( -17.20, z-score = -1.07, R) >droEre2.scaffold_4820 2612479 100 - 10470090 UUGCGGAUCGAAGCCACGUAAGUGUUCCAGAUAGUUGCAGUGGCCACAGCUUCAGUUGCGGUUUCGGUGCUCAGUAUUGAUUCCAUCCAAUC-UUUUGGAA-------------- ..((((...((((((((....))).........((.((....)).)).)))))..))))((..(((((((...)))))))..)).(((((..-..))))).-------------- ( -29.90, z-score = -1.44, R) >consensus UUGCGUAUCGAAGCCACGUAAGUGUUCCAGAUAGUUGGAGUCGCCACAGCUUCAGUUGCGGUUUCGAUGUUCAGUAUUGAUUCUGUCCAUUCGUUUUAGAA__A___________ .((.(((((((((((..((((.((........(((((.........))))).)).))))))))))))))).)).......................................... (-13.34 = -14.78 + 1.44)

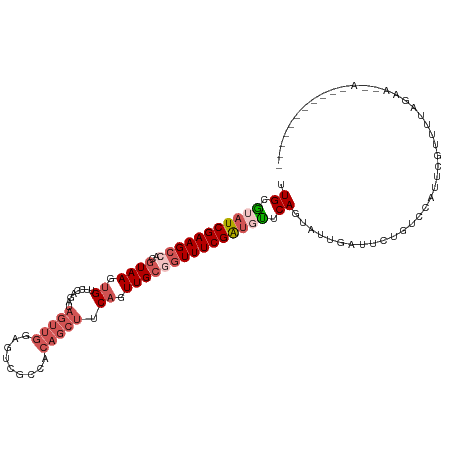

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:40:20 2011