
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,089,777 – 20,089,859 |
| Length | 82 |
| Max. P | 0.878416 |

| Location | 20,089,777 – 20,089,859 |
|---|---|
| Length | 82 |
| Sequences | 9 |
| Columns | 84 |
| Reading direction | reverse |
| Mean pairwise identity | 70.17 |
| Shannon entropy | 0.61471 |
| G+C content | 0.53306 |
| Mean single sequence MFE | -26.99 |
| Consensus MFE | -12.06 |
| Energy contribution | -12.28 |
| Covariance contribution | 0.22 |
| Combinations/Pair | 1.60 |
| Mean z-score | -1.45 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.03 |
| SVM RNA-class probability | 0.878416 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
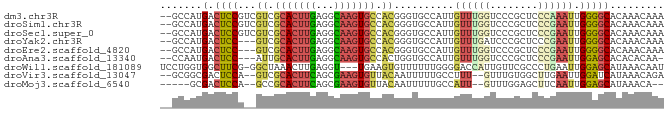
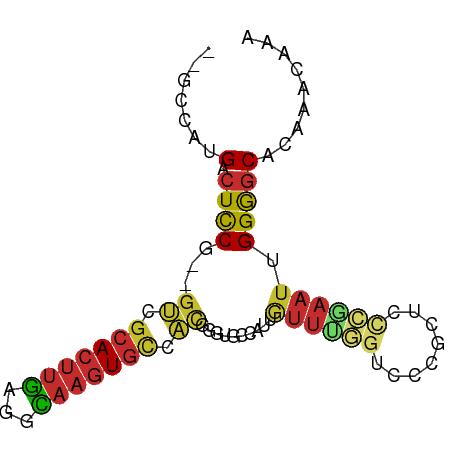
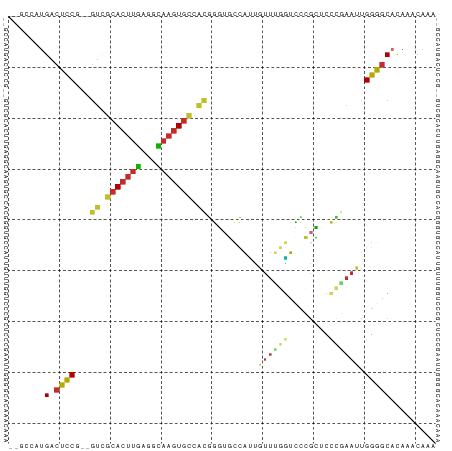
>dm3.chr3R 20089777 82 - 27905053 --GCCAUGACUCCGUCGUCGCACUUGAGGCAAGUGCCACGGGUGCCAUUGUUUGGUCCCGCUCCCAAAUUGGGGCACAAACAAA --.....(((......)))(((((((.(((....))).)))))))..(((((((((((((.........)))))).))))))). ( -32.60, z-score = -2.18, R) >droSim1.chr3R 19920792 82 - 27517382 --GCCAUGACUCCGUCGUCGCACUUGAGGCAAGUGCCACGGGUGCCAUUGUUUGGUCCCGCUCCCGAAUUGGGGCACAAACAAA --.....(((......)))(((((((.(((....))).)))))))..(((((((((((((.........)))))).))))))). ( -32.60, z-score = -1.93, R) >droSec1.super_0 20422197 82 - 21120651 --GCCAUGACUCCGUCGUCGCACUUGAGGCAAGUGCCACGGGUGCCAUUGUUUGGUCCCGCUCCCGAAUUGGGGCACAAACAAA --.....(((......)))(((((((.(((....))).)))))))..(((((((((((((.........)))))).))))))). ( -32.60, z-score = -1.93, R) >droYak2.chr3R 2415034 79 + 28832112 --GCCAUGACUCC---GUCGCACUUGAGGCAAGUGCCACGGGUGCCAUUGUUUGAUCCCGCUCCCGAAUUGGGGCACAAACAAA --......((.((---((.(((((((...))))))).))))))....(((((((.....(((((......))))).))))))). ( -27.60, z-score = -1.27, R) >droEre2.scaffold_4820 2497152 79 + 10470090 --GCCAUGACUCC---GUCGCACUUGAGGCAAGUGCCACGGGUGCCAUUGUUUGGUCCCGCUCCCGAAUUGGGGCACAAACAAA --......((.((---((.(((((((...))))))).))))))....(((((((((((((.........)))))).))))))). ( -31.20, z-score = -1.79, R) >droAna3.scaffold_13340 1290573 78 + 23697760 --CCAAUGACUCC---AUUGCACUUGAGGCAAGUGCCACUGGUGCCAUUGUUUGGUCCCGCUCCCGAAUUGGAGCACACACAA- --.(((((.(.((---(..(((((((...)))))))...))).).))))).........(((((......)))))........- ( -25.10, z-score = -1.43, R) >droWil1.scaffold_181089 3150160 80 - 12369635 UCCUGGUGGCUUCG-GGCUAAACUUGAGGU---UGAAGUGUUUUUUGGGGACCAUUGUUCGCCCUGAAUUGGAGCAUAAACAAU (((..(..((((((-(.((.......)).)---)))))).....)..)))...((((((.((((......)).))...)))))) ( -18.20, z-score = 0.82, R) >droVir3.scaffold_13047 10925515 78 - 19223366 --GCGGCGACUCCA--GUCGCACUUCAGCGAAGUGUUACAAUUUUUGCCUUU--GUUUGUGGCUUGAAUUGGAUCAUAAACAGA --..(((((.....--((.(((((((...))))))).)).....)))))(((--((((((((((......)).))))))))))) ( -24.50, z-score = -2.21, R) >droMoj3.scaffold_6540 3298024 73 + 34148556 -----GCGACUCCA--GCCGCACUUCAGCGAAGUGUUACAAUUUUUGCCAUU--GUUUGGAGCUUCAAUUGGAGCAUAAACA-- -----((((.....--...(((((((...)))))))........))))...(--(((((..(((((....))))).))))))-- ( -18.49, z-score = -1.10, R) >consensus __GCCAUGACUCCG__GUCGCACUUGAGGCAAGUGCCACGGGUGCCAUUGUUUGGUCCCGCUCCCGAAUUGGGGCACAAACAAA .......(.((((...((.(((((((...))))))).))..........((((((........)))))).)))))......... (-12.06 = -12.28 + 0.22)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:40:08 2011