
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,065,997 – 20,066,146 |
| Length | 149 |
| Max. P | 0.963894 |

| Location | 20,065,997 – 20,066,112 |
|---|---|
| Length | 115 |
| Sequences | 13 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 63.71 |
| Shannon entropy | 0.81147 |
| G+C content | 0.57895 |
| Mean single sequence MFE | -40.24 |
| Consensus MFE | -15.39 |
| Energy contribution | -14.93 |
| Covariance contribution | -0.46 |
| Combinations/Pair | 1.67 |
| Mean z-score | -1.46 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.73 |
| SVM RNA-class probability | 0.963894 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
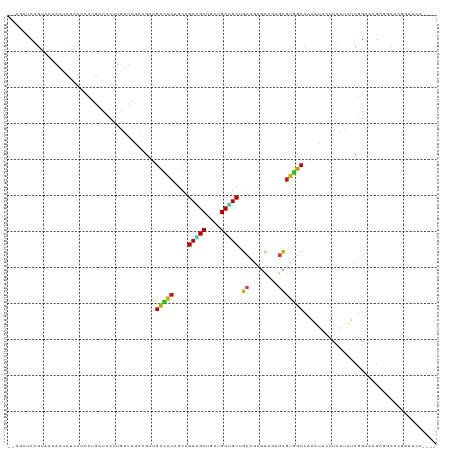
>dm3.chr3R 20065997 115 - 27905053 CAAGAUCCGAACUAAUCUCAACCUGAUCCUUGUUAACCGCUGCCACAGCAGCAGCUGCUGCUGCGGCCGAGCAGAGCGUGGCCAAGUUCUUCAUCUCGGCCCGAGGCUCCGAUCC---- (((((((.(.............).))).)))).....((..(((.(((((((....))))))).(((((((.(((((........)))))....)))))))...)))..))....---- ( -40.12, z-score = -1.68, R) >droSim1.chr3R 19897693 115 - 27517382 CAAGAUCCGAACUAAUCUCAACCUGAUCCUUGUUAACCGCUGCCACAGCAGCAGCUGCUGCUGCGGCCGAGCAGAGCGUGGCCAAGUUCUUCAUCUCGGCAGGAGGCUCGGAUCC---- ...(((((((....(((.......)))...........(((.((.(((((((....)))))))..((((((.(((((........)))))....)))))).)).)))))))))).---- ( -44.60, z-score = -2.01, R) >droSec1.super_0 20398361 115 - 21120651 CAAGAUCCGAACUAAUCUCAACCUGAUCCUUGUUAACCGCUGCCACAGCAGCAGCUGCUGCUGCGGCCGAGCACAGCGUGGCCAAGUUCUUCAUCUCGGCAGGAGGCUCGGAUCC---- ...(((((((....(((.......))).........((.(((((.(((((((....))))))).(((((.((...)).)))))..............)))))..)).))))))).---- ( -44.30, z-score = -2.28, R) >droYak2.chr3R 2391006 115 + 28832112 CAAGUCCCGAACUAAUCUCAUCCUGAUCCUUGUUAACCGCAGCCACUGCAGCAGCUGCUGCUGCGGCCGAGCAGAGCGUGGCCAAGUUCUUCAUCUCGGCGCGAGGCUCCGAUCC---- ...................(((..((.((((((....((((((....((((...)))).))))))((((((.(((((........)))))....)))))))))))).)).)))..---- ( -38.20, z-score = -0.54, R) >droEre2.scaffold_4820 2473589 115 + 10470090 CAAGUUCCGAACUAAUCUCAUCCUGAUCCUUGUUAACCGCAGCCACAGCAGCAGCUGCUGCUGCGGCCGAGCAGAGCGUGGCCAAAUUUUUCAUCUCGGCCCUAGGCUCCGAUCC---- ........................((((..((((..(((((((..((((....))))..)))))))...))))((((.(((((..............))))..).)))).)))).---- ( -37.24, z-score = -1.65, R) >droAna3.scaffold_13340 1266563 115 + 23697760 CGAUAUCAGUGCUGAUCUCGUCCUGCUCCUUGUUGGCGGCUGCCACAGCCGCUGCUGCGGCUGCUGCUGAACAGAGCGUGGCCAGGUUCUUCAUCUCGGAUCGAGGCUCGCCUCC---- .((.((((....)))).))(((.(((((..((((((((((.(((.((((....)))).))).)))))).))))))))).)))..(((((........)))))((((....)))).---- ( -47.30, z-score = -1.02, R) >dp4.chr2 14603904 117 - 30794189 --AUAUCUGUACUGAUUUCAUCUUCAUCCCUGUGGCCUGCUGCCACAGCAGCGGCCGCUGCUGCUGCUGAACAAAGUGUGGCUAGAUUCUUCAUUUCGGGACGCUGCGGGUCCUCGUCC --..(((((((.(((........)))((((..(((((((((....((((((((((....)))))))))).....)))).)))))((....)).....))))...)))))))........ ( -39.10, z-score = -1.05, R) >droPer1.super_0 6008963 117 + 11822988 --UGAUCUGUACUGAGGUCAUCUUCAUCCUUGUGGCCUGCUGCCACAGCAGCGGCCGCUGCUGCUGCUGAACAAAGUGUGGCUAGAUUCUUCAUUUCGGGACGCUGCGGGUCCUCGUCC --.((((((((.(((((....)))))((((..(((((((((....((((((((((....)))))))))).....)))).)))))((....)).....))))...))))))))....... ( -42.80, z-score = -1.03, R) >droWil1.scaffold_181089 9527349 110 + 12369635 ---CUACAUUUUCAGCAUUGUCCUUUUUCUUGUUAGCAGCAGCUUCAGCAGCCGCCACAGCGGCCGCUGAGCACAAUGUAGCCAGAUUCUUCAUUUCGGCUGGGUCGAUACCC------ ---...........(((((((.........(((.....)))(((.((((.(((((....))))).))))))))))))))((((..(........)..))))((((....))))------ ( -36.20, z-score = -2.98, R) >droMoj3.scaffold_6540 7516705 107 - 34148556 --AUGUCGGUGACAAUUUCAUCGAGUUCCUUAUUUGUCGCUGUUACCUGGGCGGCCAUGGCUGCUGCGGAGCAGAGUGUUGCCAGAUUCUUCAUUUCAGCCCGGUUCUC---------- --....(((((((((..................)))))))))....((((((((((((..(((((....))))).)))..))).((....))......)))))).....---------- ( -34.17, z-score = -0.92, R) >droGri2.scaffold_14906 12061179 107 + 14172833 --UUAUCAGUUGCAACUUCUUCGGACUCCUUAUUAGCUGCUGUUACGGCAGCCGCCAUGGCUGCUGCCGAGCACAGCGUGGCCAAAUUCUUCAUCUCUGCACGCGUUUC---------- --......((..(....((....))..........((((.((((.(((((((.((....)).))))))))))))))))..))...........................---------- ( -35.30, z-score = -1.97, R) >droVir3.scaffold_12855 9142308 107 + 10161210 --AUAUCGGUAGCAAUUUCAUCGAUUUCCUUAUUUGCCGCUGUCACCUGAGCCGCCAUGGCGGCUGCAGAGCAGAGCGUGGCCAGAUUCUUCAUUUCGGCCCGGGUUUC---------- --..((((((.........))))))..........(((.((((...(((((((((....)))))).))).))))..((.((((..(........)..)))))))))...---------- ( -33.80, z-score = -0.48, R) >anoGam1.chr3R 11130360 94 - 53272125 ------CGCUUCCGGAGCCGCUUCCCUGUCCGGCGGCGGCCGCCGCAGCAGCUGCUGCUGCAGCCGAAAUGUCCAGGCCGGCGGAGCGCGUGAGCGAUUC------------------- ------((((..(((..(((((..((((..((....((((.((.(((((....))))).)).))))...))..))))..)))))..).))..))))....------------------- ( -50.00, z-score = -1.37, R) >consensus __AGAUCCGUACUAAUCUCAUCCUGAUCCUUGUUAGCCGCUGCCACAGCAGCAGCUGCUGCUGCUGCCGAGCAGAGCGUGGCCAAAUUCUUCAUCUCGGCACGAGGCUCGGAUCC____ .........................................(((((....(((((....))))).((........)))))))..................................... (-15.39 = -14.93 + -0.46)


| Location | 20,066,033 – 20,066,146 |
|---|---|
| Length | 113 |
| Sequences | 13 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 59.60 |
| Shannon entropy | 0.89271 |
| G+C content | 0.57435 |
| Mean single sequence MFE | -39.26 |
| Consensus MFE | -10.98 |
| Energy contribution | -10.70 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.75 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.28 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.820080 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
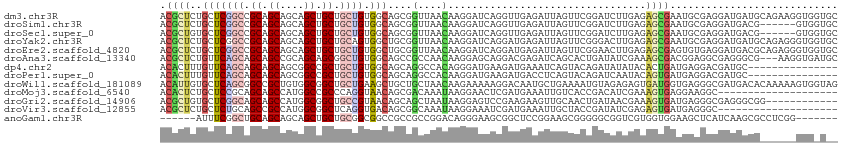
>dm3.chr3R 20066033 113 + 27905053 ACGCUCUGCUCGGCCGCAGCAGCAGCUGCUGCUGUGGCAGCGGUUAACAAGGAUCAGGUUGAGAUUAGUUCGGAUCUUGAGAGCGAAUGCGAGGAUGAUGCAGAAGGUGGUGC .((((((((((((((((((((((....))))))))))).((((((......))))...(..(((((......)))))..)..))...........))).)))))..))).... ( -43.70, z-score = -1.28, R) >droSim1.chr3R 19897729 107 + 27517382 ACGCUCUGCUCGGCCGCAGCAGCAGCUGCUGCUGUGGCAGCGGUUAACAAGGAUCAGGUUGAGAUUAGUUCGGAUCUUGAGAGCGAAUGCGAGGAUGACG------GUGGUGC .(((..(((((.(((((((((((....)))))))))))...((((......))))...(..(((((......)))))..))))))...))).........------....... ( -41.90, z-score = -1.15, R) >droSec1.super_0 20398397 107 + 21120651 ACGCUGUGCUCGGCCGCAGCAGCAGCUGCUGCUGUGGCAGCGGUUAACAAGGAUCAGGUUGAGAUUAGUUCGGAUCUUGAGAGCGAAUGCGAGGAUGACG------GUGGUGC .((((((.(((((((((((((((....))))))))))).((((((......))))...(..(((((......)))))..)..)).....))))....)))------))).... ( -44.40, z-score = -2.13, R) >droYak2.chr3R 2391042 113 - 28832112 ACGCUCUGCUCGGCCGCAGCAGCAGCUGCUGCAGUGGCUGCGGUUAACAAGGAUCAGGAUGAGAUUAGUUCGGGACUUGAGAGCGAAUGCGAGGAUGAUGCAGAGGGUGGUGC ((.(((((((((.((...((((((((..(....)..))))).(((..((((..((.((((.......))))))..))))..)))...)))..)).))).)))))).))..... ( -44.40, z-score = -1.68, R) >droEre2.scaffold_4820 2473625 113 - 10470090 ACGCUCUGCUCGGCCGCAGCAGCAGCUGCUGCUGUGGCUGCGGUUAACAAGGAUCAGGAUGAGAUUAGUUCGGAACUUGAGAGCGAGUGUGAGGAUGACGCAGAGGGUGGUGC .(((((((((((((((((((.(((((....))))).)))))))))..((((..((.((((.......)))).)).)))).))))..((((.......))))..)))))).... ( -48.70, z-score = -2.76, R) >droAna3.scaffold_13340 1266599 110 - 23697760 ACGCUCUGUUCAGCAGCAGCCGCAGCAGCGGCUGUGGCAGCCGCCAACAAGGAGCAGGACGAGAUCAGCACUGAUAUCGAAAGCGACGGAGGCGAGGGCG---AAGGUGAUGC .((((((.....((.((((((((....)))))))).))...((((.....(..((....(((.((((....)))).)))...))..)...))))))))))---.......... ( -43.40, z-score = -1.26, R) >dp4.chr2 14603944 98 + 30794189 ACACUUUGUUCAGCAGCAGCAGCGGCCGCUGCUGUGGCAGCAGGCCACAGGGAUGAAGAUGAAAUCAGUACAGAUAUAUACACUGAUGAGGACGAUGC--------------- .(((((..(((((((((.((....)).))))))(((((.....)))))..)))..))).))..((((((............))))))...........--------------- ( -32.40, z-score = -1.75, R) >droPer1.super_0 6009003 98 - 11822988 ACACUUUGUUCAGCAGCAGCAGCGGCCGCUGCUGUGGCAGCAGGCCACAAGGAUGAAGAUGACCUCAGUACAGAUCAAUACAGUGAUGAGGACGAUGC--------------- .(((((..(((((((((.((....)).))))))(((((.....)))))..)))..))).)).(((((.(((...........))).))))).......--------------- ( -36.50, z-score = -2.35, R) >droWil1.scaffold_181089 9527383 113 - 12369635 ACAUUGUGCUCAGCGGCCGCUGUGGCGGCUGCUGAAGCUGCUGCUAACAAGAAAAAGGACAAUGCUGAAAAUGUAGAGAGUGAUGGUGAGGGCGAUGACACAAAAAGUGGUAG .((((((.((((.((..(((((..((((((.....))))))..)...............(..(((.......)))..))))).)).)))).)))))).(((.....))).... ( -36.80, z-score = -2.11, R) >droMoj3.scaffold_6540 7516735 93 + 34148556 ACACUCUGCUCCGCAGCAGCCAUGGCCGCCCAGGUAACAGCGACAAAUAAGGAACUCGAUGAAAUUGUCACCGACAUCGAAAGUGAGGAAGGC-------------------- .((((.((((((...((.((....)).))...))....)))).............((((((...(((....))))))))).))))........-------------------- ( -17.80, z-score = 1.61, R) >droGri2.scaffold_14906 12061209 101 - 14172833 ACGCUGUGCUCGGCAGCAGCCAUGGCGGCUGCCGUAACAGCAGCUAAUAAGGAGUCCGAAGAAGUUGCAACUGAUAACGAAAGUGAUGAGGGCGAGGGCGG------------ .((((.(((((((((((.((....)).))))))).....((((((.....((...)).....))))))..((.((.((....)).)).))))))..)))).------------ ( -40.60, z-score = -2.61, R) >droVir3.scaffold_12855 9142338 93 - 10161210 ACGCUCUGCUCUGCAGCCGCCAUGGCGGCUCAGGUGACAGCGGCAAAUAAGGAAAUCGAUGAAAUUGCUACCGAUAUCGAGAGUGAUGAGGGC-------------------- .((((((...((((((((((....))))))..(....).))))........((.((((.............)))).)).))))))........-------------------- ( -29.32, z-score = -0.39, R) >anoGam1.chr3R 11130390 100 + 53272125 ------AUUUCGGCUGCAGCAGCAGCUGCUGCGGCGGCCGCCGCCGGACAGGGAAGCGGCUCCGGAAGCGGGGGCGGUCGUGGUGGAAGCUCAUCAAGCGCCUCGG------- ------.(((((((((((((((...))))))))))((((((..((......))..)))))).)))))...((((((....((((((....))))))..))))))..------- ( -50.40, z-score = -0.61, R) >consensus ACGCUCUGCUCGGCAGCAGCAGCAGCUGCUGCUGUGGCAGCGGCUAACAAGGAUCAGGAUGAGAUUAGUACGGAUCUUGAGAGUGAAGGCGACGAUGACG______GUG_U__ .((((..(((((((.((.((....)).)).)))).)))....(....).................................))))............................ (-10.98 = -10.70 + -0.28)

| Location | 20,066,033 – 20,066,146 |
|---|---|
| Length | 113 |
| Sequences | 13 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 59.60 |
| Shannon entropy | 0.89271 |
| G+C content | 0.57435 |
| Mean single sequence MFE | -32.15 |
| Consensus MFE | -10.83 |
| Energy contribution | -11.28 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.54 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.34 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.43 |
| SVM RNA-class probability | 0.939739 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
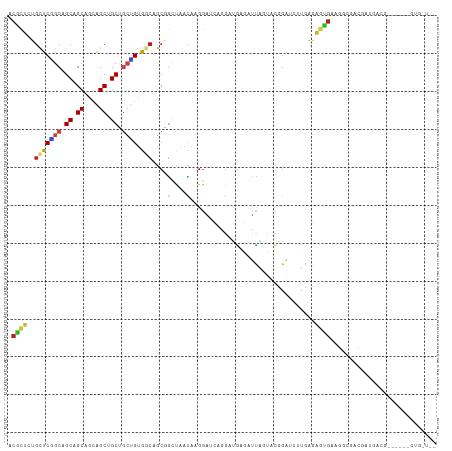
>dm3.chr3R 20066033 113 - 27905053 GCACCACCUUCUGCAUCAUCCUCGCAUUCGCUCUCAAGAUCCGAACUAAUCUCAACCUGAUCCUUGUUAACCGCUGCCACAGCAGCAGCUGCUGCUGCGGCCGAGCAGAGCGU (((........)))..............(((((((((((((.(.............).))).))))......((((((.(((((((....))))))).)))..))))))))). ( -35.12, z-score = -2.30, R) >droSim1.chr3R 19897729 107 - 27517382 GCACCAC------CGUCAUCCUCGCAUUCGCUCUCAAGAUCCGAACUAAUCUCAACCUGAUCCUUGUUAACCGCUGCCACAGCAGCAGCUGCUGCUGCGGCCGAGCAGAGCGU .......------...............(((((((((((((.(.............).))).))))......((((((.(((((((....))))))).)))..))))))))). ( -33.32, z-score = -2.02, R) >droSec1.super_0 20398397 107 - 21120651 GCACCAC------CGUCAUCCUCGCAUUCGCUCUCAAGAUCCGAACUAAUCUCAACCUGAUCCUUGUUAACCGCUGCCACAGCAGCAGCUGCUGCUGCGGCCGAGCACAGCGU .......------.........(((....((((.(((((((.(.............).))).)))).........(((.(((((((....))))))).))).))))...))). ( -30.12, z-score = -1.48, R) >droYak2.chr3R 2391042 113 + 28832112 GCACCACCCUCUGCAUCAUCCUCGCAUUCGCUCUCAAGUCCCGAACUAAUCUCAUCCUGAUCCUUGUUAACCGCAGCCACUGCAGCAGCUGCUGCUGCGGCCGAGCAGAGCGU .....((.((((((.........((....))..........(((((..(((.......)))....)))..(((((((....((((...)))).))))))).)).)))))).)) ( -29.50, z-score = -0.63, R) >droEre2.scaffold_4820 2473625 113 + 10470090 GCACCACCCUCUGCGUCAUCCUCACACUCGCUCUCAAGUUCCGAACUAAUCUCAUCCUGAUCCUUGUUAACCGCAGCCACAGCAGCAGCUGCUGCUGCGGCCGAGCAGAGCGU (((........)))..............((((((...((((..(((..(((.......)))....)))..(((((((..((((....))))..)))))))..)))))))))). ( -33.40, z-score = -1.91, R) >droAna3.scaffold_13340 1266599 110 + 23697760 GCAUCACCUU---CGCCCUCGCCUCCGUCGCUUUCGAUAUCAGUGCUGAUCUCGUCCUGCUCCUUGUUGGCGGCUGCCACAGCCGCUGCUGCGGCUGCUGCUGAACAGAGCGU ..........---.((...((....))..))...(((.((((....)))).)))....((((..((((((((((.(((.((((....)))).))).)))))).)))))))).. ( -36.40, z-score = -0.09, R) >dp4.chr2 14603944 98 - 30794189 ---------------GCAUCGUCCUCAUCAGUGUAUAUAUCUGUACUGAUUUCAUCUUCAUCCCUGUGGCCUGCUGCCACAGCAGCGGCCGCUGCUGCUGCUGAACAAAGUGU ---------------((((.......(((((((((......)))))))))((((..........((((((.....))))))((((((((....))))))))))))....)))) ( -30.60, z-score = -1.64, R) >droPer1.super_0 6009003 98 + 11822988 ---------------GCAUCGUCCUCAUCACUGUAUUGAUCUGUACUGAGGUCAUCUUCAUCCUUGUGGCCUGCUGCCACAGCAGCGGCCGCUGCUGCUGCUGAACAAAGUGU ---------------((((.(.(((((.....((((......))))))))).)...((((....((((((.....))))))((((((((....))))))))))))....)))) ( -33.60, z-score = -1.63, R) >droWil1.scaffold_181089 9527383 113 + 12369635 CUACCACUUUUUGUGUCAUCGCCCUCACCAUCACUCUCUACAUUUUCAGCAUUGUCCUUUUUCUUGUUAGCAGCAGCUUCAGCAGCCGCCACAGCGGCCGCUGAGCACAAUGU ....(((.....))).................................(((((((.........(((.....)))(((.((((.(((((....))))).)))))))))))))) ( -28.80, z-score = -2.87, R) >droMoj3.scaffold_6540 7516735 93 - 34148556 --------------------GCCUUCCUCACUUUCGAUGUCGGUGACAAUUUCAUCGAGUUCCUUAUUUGUCGCUGUUACCUGGGCGGCCAUGGCUGCUGCGGAGCAGAGUGU --------------------........((((...((..(((((((.....)))))))..))...........(((((.((..((((((....))))))..))))))))))). ( -29.00, z-score = -0.56, R) >droGri2.scaffold_14906 12061209 101 + 14172833 ------------CCGCCCUCGCCCUCAUCACUUUCGUUAUCAGUUGCAACUUCUUCGGACUCCUUAUUAGCUGCUGUUACGGCAGCCGCCAUGGCUGCUGCCGAGCACAGCGU ------------........((.((.((.((....)).)).))..))....((....))..........((((.((((.(((((((.((....)).))))))))))))))).. ( -31.50, z-score = -1.84, R) >droVir3.scaffold_12855 9142338 93 + 10161210 --------------------GCCCUCAUCACUCUCGAUAUCGGUAGCAAUUUCAUCGAUUUCCUUAUUUGCCGCUGUCACCUGAGCCGCCAUGGCGGCUGCAGAGCAGAGCGU --------------------.........((.(((((((.((((((.....................)))))).))))..(((((((((....)))))).)))....))).)) ( -28.30, z-score = -1.00, R) >anoGam1.chr3R 11130390 100 - 53272125 -------CCGAGGCGCUUGAUGAGCUUCCACCACGACCGCCCCCGCUUCCGGAGCCGCUUCCCUGUCCGGCGGCGGCCGCCGCAGCAGCUGCUGCUGCAGCCGAAAU------ -------....((((.(((.((.(......)))))).)))).(((....))).((((((.........))))))(((.((.(((((....))))).)).))).....------ ( -38.30, z-score = 0.31, R) >consensus __A_CAC______CGUCAUCGUCCCCAUCACUCUCAAGAUCCGUACUAAUCUCAUCCUGAUCCUUGUUAGCCGCUGCCACAGCAGCAGCUGCUGCUGCUGCCGAGCAGAGCGU ............................((((........................................((((...)))).(((((....)))))..........)))). (-10.83 = -11.28 + 0.45)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:40:07 2011