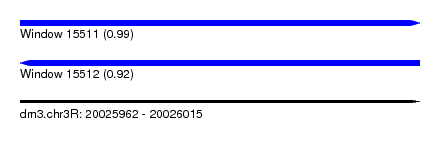

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 20,025,962 – 20,026,015 |
| Length | 53 |
| Max. P | 0.992820 |

| Location | 20,025,962 – 20,026,015 |
|---|---|
| Length | 53 |
| Sequences | 6 |
| Columns | 61 |
| Reading direction | forward |
| Mean pairwise identity | 72.50 |
| Shannon entropy | 0.49995 |
| G+C content | 0.55270 |
| Mean single sequence MFE | -18.87 |
| Consensus MFE | -9.32 |
| Energy contribution | -9.10 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.53 |
| Mean z-score | -2.66 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.57 |
| SVM RNA-class probability | 0.992820 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
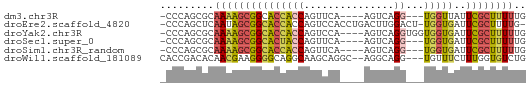
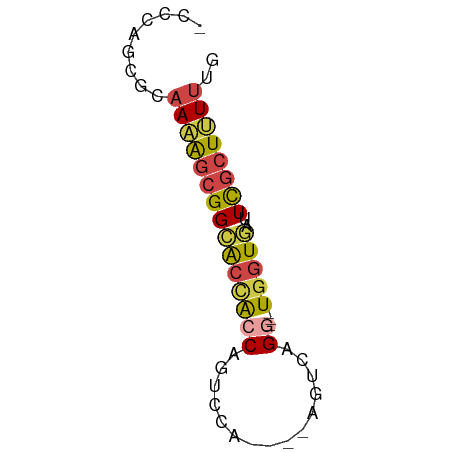
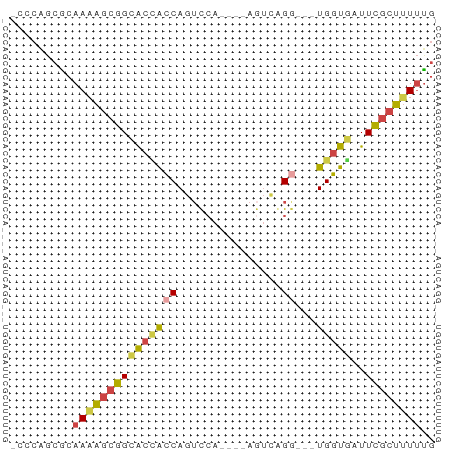
>dm3.chr3R 20025962 53 + 27905053 -CCCAGCGCAAAAGCGGCACCACCAGUUCA----AGUCAGG---UGGUUAUUCGCUUUUUG -........((((((((.((((((......----.....))---))))...)))))))).. ( -15.90, z-score = -2.62, R) >droEre2.scaffold_4820 2438476 58 - 10470090 -CCCAGCUCAAUAGCGGCACCACCAGUCCACCUGACUUGGACU-UGGUGAUUCGCUUUUG- -.......(((.(((((...((((((((((.......))))).-)))))..))))).)))- ( -20.30, z-score = -3.38, R) >droYak2.chr3R 2355428 56 - 28832112 -CCCAGCGCAAAAGCGGCACCACCAGUCCA----AGUCAGGUGGUGGUGAUUCGCUUUUUG -........(((((((.((((((((..((.----.....))))))))))...))))))).. ( -22.30, z-score = -3.14, R) >droSec1.super_0 20363649 53 + 21120651 -CCCAGCGCAAAAGCGGCACUACCAGUUCA----AGUCAGG---UGGUGAUUCGCUUUUUG -........(((((((.(((((((......----.....))---)))))...))))))).. ( -16.70, z-score = -2.12, R) >droSim1.chr3R_random 970166 53 + 1307089 -CCCAGCGCAAAAGCGGCACCACCAGUUCA----AGUCAGG---UGGUGAUUCGCUUUUUG -........(((((((.(((((((......----.....))---)))))...))))))).. ( -19.00, z-score = -3.01, R) >droWil1.scaffold_181089 9493969 56 - 12369635 CACCGACACAACGAAGGGGCAGGCAAGCAGGC--AGGCAGG---UGUUUCUUUGGUGUCUG ....(((((..(((((..(((.((..(....)--..))...---)))..)))))))))).. ( -19.00, z-score = -1.68, R) >consensus _CCCAGCGCAAAAGCGGCACCACCAGUCCA____AGUCAGG___UGGUGAUUCGCUUUUUG .........((((((((...(((((..((..........))...)))))..)))))))).. ( -9.32 = -9.10 + -0.22)

| Location | 20,025,962 – 20,026,015 |
|---|---|
| Length | 53 |
| Sequences | 6 |
| Columns | 61 |
| Reading direction | reverse |
| Mean pairwise identity | 72.50 |
| Shannon entropy | 0.49995 |
| G+C content | 0.55270 |
| Mean single sequence MFE | -17.28 |
| Consensus MFE | -8.72 |
| Energy contribution | -10.03 |
| Covariance contribution | 1.31 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.921670 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
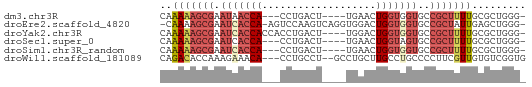
>dm3.chr3R 20025962 53 - 27905053 CAAAAAGCGAAUAACCA---CCUGACU----UGAACUGGUGGUGCCGCUUUUGCGCUGGG- ..(((((((....((((---((.....----......))))))..)))))))........- ( -17.00, z-score = -2.41, R) >droEre2.scaffold_4820 2438476 58 + 10470090 -CAAAAGCGAAUCACCA-AGUCCAAGUCAGGUGGACUGGUGGUGCCGCUAUUGAGCUGGG- -(((.((((.(((((((-.(((((.......))))))))))))..)))).))).......- ( -24.20, z-score = -3.37, R) >droYak2.chr3R 2355428 56 + 28832112 CAAAAAGCGAAUCACCACCACCUGACU----UGGACUGGUGGUGCCGCUUUUGCGCUGGG- ..(((((((...((((((((((.....----.))..)))))))).)))))))........- ( -22.30, z-score = -3.17, R) >droSec1.super_0 20363649 53 - 21120651 CAAAAAGCGAAUCACCA---CCUGACU----UGAACUGGUAGUGCCGCUUUUGCGCUGGG- ..(((((((...(((.(---((.....----......))).))).)))))))........- ( -12.50, z-score = -0.49, R) >droSim1.chr3R_random 970166 53 - 1307089 CAAAAAGCGAAUCACCA---CCUGACU----UGAACUGGUGGUGCCGCUUUUGCGCUGGG- ..(((((((...(((((---((.....----......))))))).)))))))........- ( -19.00, z-score = -2.78, R) >droWil1.scaffold_181089 9493969 56 + 12369635 CAGACACCAAAGAAACA---CCUGCCU--GCCUGCUUGCCUGCCCCUUCGUUGUGUCGGUG ..(((((....(((...---...((..--((......))..))...)))...))))).... ( -8.70, z-score = 0.80, R) >consensus CAAAAAGCGAAUCACCA___CCUGACU____UGAACUGGUGGUGCCGCUUUUGCGCUGGG_ ..(((((((.(((((((...................)))))))..)))))))......... ( -8.72 = -10.03 + 1.31)
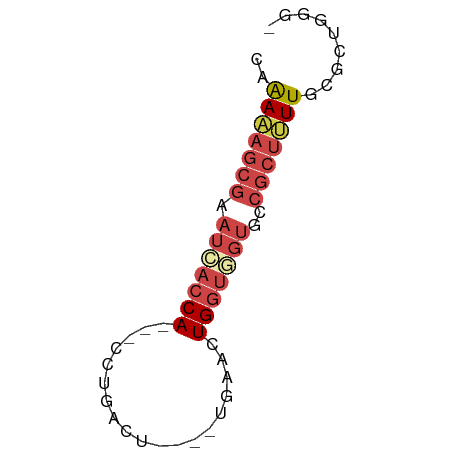
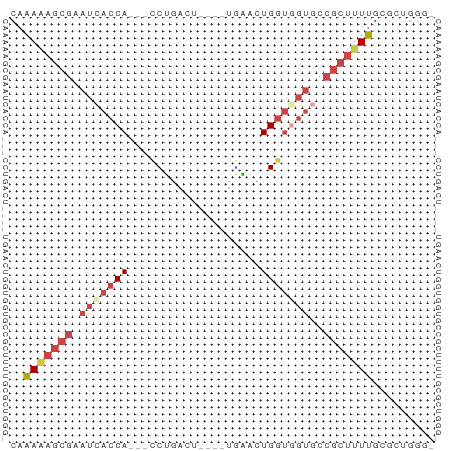
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:40:00 2011