
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 19,948,151 – 19,948,351 |
| Length | 200 |
| Max. P | 0.750932 |

| Location | 19,948,151 – 19,948,249 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.42 |
| Shannon entropy | 0.28351 |
| G+C content | 0.48047 |
| Mean single sequence MFE | -34.18 |
| Consensus MFE | -23.15 |
| Energy contribution | -24.27 |
| Covariance contribution | 1.11 |
| Combinations/Pair | 1.13 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.750932 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
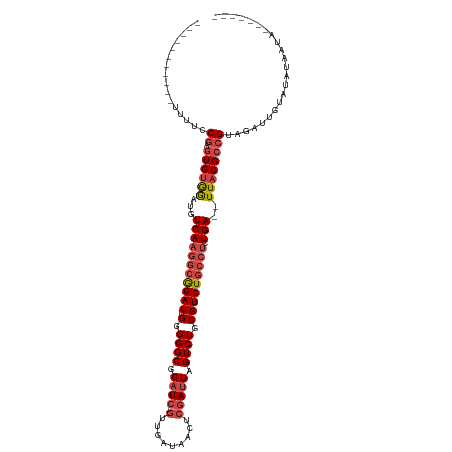
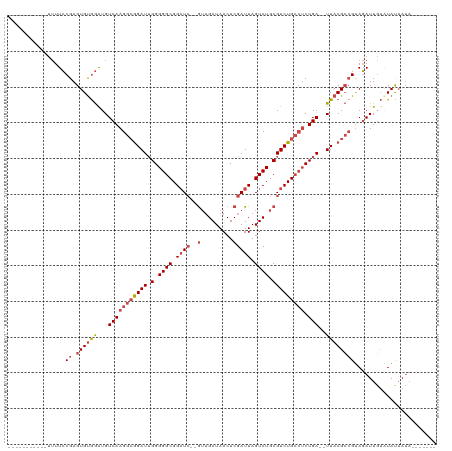
>dm3.chr3R 19948151 98 + 27905053 -----------UUUUCCGAGUGUGGAUGUCAAGGCGGAUUGGGGGCGGAUC--GUUGAUAACUCGAUUAGUCUGCGUCUGCCUCUGA--UUACGCCGUAGAUUGUAUAUAAUA------- -----------........(((((.(.(((..(((((((..((((((((.(--((.(((..........))).)))))))))))..)--)).))))...)))).)))))....------- ( -34.00, z-score = -2.35, R) >droSim1.chr3R 19779783 98 + 27517382 -----------UUUUCCGAGUGUGGAUGUCAAGGCGGAUUGGGGGCGGAUC--GUUGAUAACUCGAUUAGUCUGCGUCUGCCUCUGA--UUACGCAGUAGAUUGUAUACAAUA------- -----------........(((((.(.((((..((((((..((((((((.(--((.(((..........))).)))))))))))..)--)).)))..).)))).)))))....------- ( -32.60, z-score = -2.02, R) >droSec1.super_0 20286194 98 + 21120651 -----------UUUUCCGAGUGUGGAUGUCAAGGCGGAUUGGGGGCGGAUC--GUUGAUAACUCGAUUAGUCUGCGUCUGCCUCUGA--UUACGCCGUAGAUUGUAUACAAUA------- -----------........(((((.(.(((..(((((((..((((((((.(--((.(((..........))).)))))))))))..)--)).))))...)))).)))))....------- ( -36.20, z-score = -2.90, R) >droYak2.chr3R 2277225 105 - 28832112 -----------UUUUCCGAGUGUGGAUGUCAAGGCGGAUUGGGGGCGGAUC--GUUGAUAACUCGAUUAGUCUGCGUCUGCCGCUGA--UUACGCCGCAGAUUGUCUAUAAUAAUUAUAA -----------.........((((((((((..(((((((..(.((((((.(--((.(((..........))).))))))))).)..)--)).))))...)).)))))))).......... ( -35.30, z-score = -2.04, R) >droEre2.scaffold_4820 2359067 98 - 10470090 -----------UUUUCCGAGUGUGGAUGUCAAGGCGGAUUGGGGGCGGAUC--GUUGAUAACUCGAUUAGUCUGCGUCUGCCUCUGA--UUACGCCGUAGAUUGUGUAUAAUA------- -----------...((((....)))).(((..(((((((..((((((((.(--((.(((..........))).)))))))))))..)--)).))))...)))...........------- ( -33.40, z-score = -2.08, R) >droAna3.scaffold_13340 1149398 110 - 23697760 CACCGACUCGCCAUUGCGAGUGCAAGUGUCAAGGAAGAU-GGGGGCGGCUGCGAGAGAUAACUCGAUUAGUCUGCGUCUGGGUCUGAGAUUACGGCGUACAUUUCAUAUUA--------- (.(((((((((....))))))...(((.(((.((.((((-(.((((.....((((......))))....)))).)))))...))))).))).))).)..............--------- ( -33.60, z-score = -0.27, R) >consensus ___________UUUUCCGAGUGUGGAUGUCAAGGCGGAUUGGGGGCGGAUC__GUUGAUAACUCGAUUAGUCUGCGUCUGCCUCUGA__UUACGCCGUAGAUUGUAUAUAAUA_______ ................((.((((((...(((((((((((.....((((((...(((((....)))))..)))))))))))))).)))..))))))))....................... (-23.15 = -24.27 + 1.11)


| Location | 19,948,249 – 19,948,351 |
|---|---|
| Length | 102 |
| Sequences | 7 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.42 |
| Shannon entropy | 0.53023 |
| G+C content | 0.43115 |
| Mean single sequence MFE | -25.09 |
| Consensus MFE | -8.79 |
| Energy contribution | -11.59 |
| Covariance contribution | 2.80 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.26 |
| SVM RNA-class probability | 0.616175 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
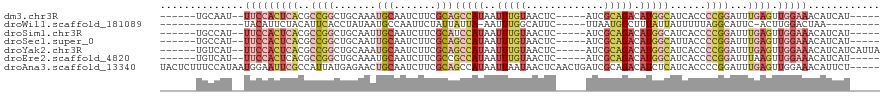
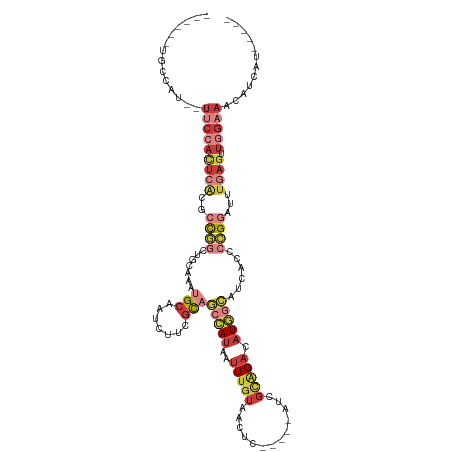
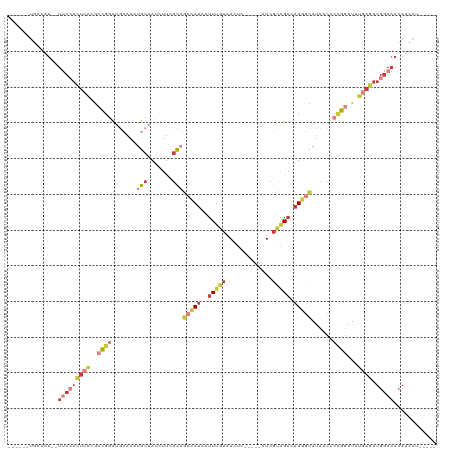
>dm3.chr3R 19948249 102 - 27905053 ------UGCAAU--UUCCACUCACGCCGGCUGCAAAUGCAAUCUUCGCAGCCAUAAUUUGUAACUC-----AUCGCAGACAUGGCAUCACCCCGGAUUUGAGUUGGAAACAUCAU----- ------.....(--(((((((((..((((.(((....))).........(((((..(((((.....-----...))))).)))))......))))...)))).))))))......----- ( -29.40, z-score = -2.40, R) >droWil1.scaffold_181089 6409481 90 + 12369635 --------------UACAUUCUACAUUCACCUAUAAUGCCAAUUCUAUUAUUA-AAUUUGGCAUUC-----UUAAUGGCUUUAUUAUUUUUAGGCAUUC-ACUUGGACUAA--------- --------------...............((...((((((((...........-...)))))))).-----..((((.((..(.....)..)).)))).-....)).....--------- ( -11.84, z-score = -1.42, R) >droSim1.chr3R 19779881 102 - 27517382 ------UGCCAU--UUCCACUCACGCCGGCUGCAAUUGCAAUCUUCGCAUCCAUAAUUUGUAACUC-----AUCGCAGACAUGGCAUCACCCCGGAUUUGAGUUGGAAACAUCAU----- ------.....(--(((((((((..((((.(((....)))..........((((..(((((.....-----...))))).)))).......))))...)))).))))))......----- ( -24.60, z-score = -1.15, R) >droSec1.super_0 20286292 102 - 21120651 ------UGCCAU--UUCCACUCACGCCGGCUGCAAUUGCAAUCUUCGCAGCCAUAAUUUGUAACUC-----AUCGCAGACAUGGCAUUACCCCGGAUUUGAGUUGGAAACAUCAU----- ------.....(--(((((((((..((((.(((....))).........(((((..(((((.....-----...))))).)))))......))))...)))).))))))......----- ( -29.40, z-score = -2.47, R) >droYak2.chr3R 2277330 107 + 28832112 ------UGUCAU--UUCCACUCACGCCGGCUGCAAAUGCAAUCUUCGCAGCCAUAAUUUGUAACUC-----AUCGCAGACAUGGCAUCACCCCGGAUUUGAGUUGGAAACAUCAUCAUUA ------.....(--(((((((((..((((.(((....))).........(((((..(((((.....-----...))))).)))))......))))...)))).))))))........... ( -29.40, z-score = -2.46, R) >droEre2.scaffold_4820 2359165 102 + 10470090 ------UGUCAU--UUCCACUCACGCCGGCUGCAAAUGCAAUCUUCGCCGCCAUAAUUUGUAACUC-----AUCGCAGACAUGGCAUCACCCCGGAUUUAAGUUGGAAACAUCAU----- ------.....(--(((((((....((((.(((....))).........(((((..(((((.....-----...))))).)))))......)))).....)).))))))......----- ( -24.40, z-score = -1.35, R) >droAna3.scaffold_13340 1149508 115 + 23697760 UACUCUUUCCAUAAUGGAAUUCGCCAUUAUGAGAACUGCAAUCUUCGCAGCCAUAAUUAAUAACUCAACUGAUCGCAGACAUCUCAUCACCCCGGAUUUGAGUUGGAAACAUUCU----- .((((..((((((((((......)))))))((((.((((.......))))..................(((....)))...))))........)))...))))((....))....----- ( -26.60, z-score = -2.71, R) >consensus ______UGCCAU__UUCCACUCACGCCGGCUGCAAAUGCAAUCUUCGCAGCCAUAAUUUGUAACUC_____AUCGCAGACAUGGCAUCACCCCGGAUUUGAGUUGGAAACAUCAU_____ ..............(((((((((..((((.......(((.......)))(((((..(((((.............))))).)))))......))))...)))).)))))............ ( -8.79 = -11.59 + 2.80)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:39:45 2011