
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 19,803,263 – 19,803,353 |
| Length | 90 |
| Max. P | 0.899808 |

| Location | 19,803,263 – 19,803,353 |
|---|---|
| Length | 90 |
| Sequences | 15 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 68.03 |
| Shannon entropy | 0.67280 |
| G+C content | 0.46721 |
| Mean single sequence MFE | -23.47 |
| Consensus MFE | -13.75 |
| Energy contribution | -12.80 |
| Covariance contribution | -0.95 |
| Combinations/Pair | 1.67 |
| Mean z-score | -0.66 |
| Structure conservation index | 0.59 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.15 |
| SVM RNA-class probability | 0.899808 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
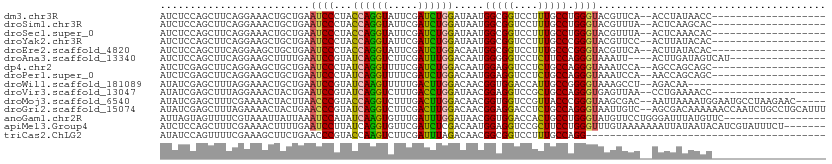
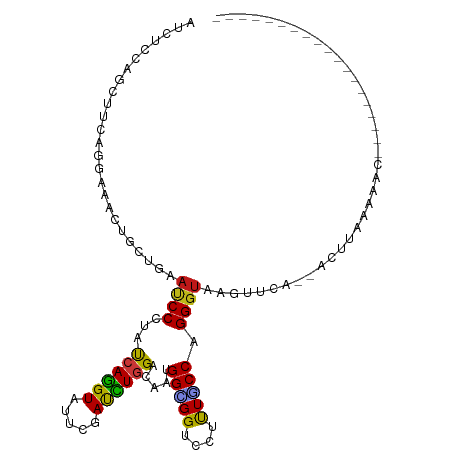
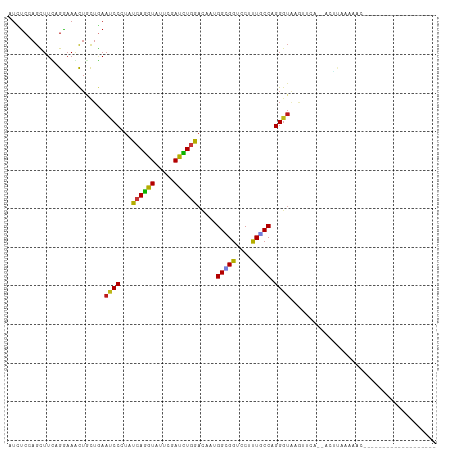
>dm3.chr3R 19803263 90 + 27905053 AUCUCCAGCUUCAGGAAACUGCUGAAUCCCUACCAGGUAUUCGAUCUGGAUAAUGGCGGUCCUUUGCCUGGGUACGUUCA--ACCUAUAACC------------------- ...(((((...(((....))).(((((.((.....)).)))))..)))))....(((((....)))))(((((.......--))))).....------------------- ( -23.70, z-score = -0.83, R) >droSim1.chr3R 19627496 90 + 27517382 AUCUCCAGCUUCAGGAAACUGCUGAAUCCCUACCAGGUAUUCGAUCUGGAUAAUGGCGGUCCUUUGCCUGGGUACGUUUA--ACUCAAGCAC------------------- .......(((((((....))).((((((((..((((((.....)))))).....(((((....))))).)))...)))))--....))))..------------------- ( -24.90, z-score = -1.04, R) >droSec1.super_0 20144778 90 + 21120651 AUCUCCAGCUUCAGGAAACUGCUGAAUCCCUACCAGGUAUUCGAUCUGGAUAAUGGCGGUCCUUUGCCUGGGUACGUUUA--ACUCAAACAC------------------- .....((((...((....))))))...(((..((((((.....)))))).....(((((....))))).)))...((((.--....))))..------------------- ( -24.10, z-score = -1.26, R) >droYak2.chr3R 2133356 90 - 28832112 AUCUCCAGCUUCAGGAAGCUGCUGAAUCCCUACCAGGUAUUCGAUCUGGAUAAUGGCGGUCCUUUGCCCGGGUACGUUCC--ACUUAUACAC------------------- .((..((((((....))))))..))..(((..((((((.....)))))).....(((((....))))).)))........--..........------------------- ( -25.00, z-score = -0.77, R) >droEre2.scaffold_4820 2215984 90 - 10470090 AUCUCCAGCUUCAGGAAGCUGCUGAAUCCCUACCAGGUAUUCGAUCUGGACAAUGGCGGUCCUUUGCCCGGGUACGUUCA--ACUUAUACAC------------------- .....((((((....)))))).((((((((..((((((.....)))))).....(((((....))))).)))...)))))--..........------------------- ( -25.80, z-score = -1.16, R) >droAna3.scaffold_13340 15941411 91 - 23697760 AUCUCCAGCUUCAGGAAGCUUUUGAAUCCGUAUCAGGUCUUCGAUUUGGACAAUGGGGGUCCUCUUCCAGGGUAAAUU----ACUUGAUAGUCAU---------------- .((...(((((....)))))...)).....((((((((....(((((..((..((((((....))))))..)))))))----)))))))).....---------------- ( -21.00, z-score = 0.52, R) >dp4.chr2 14309134 90 + 30794189 AUCUCGAGCUUCAGGAAGCUGCUGAAUCCCUAUCAGGUUUUCGAUCUGGACAAUGGAGGUCCUCUGCCAGGGUAAAUCCA--AGCCAGCAGC------------------- .((..(.....)..)).(((((((((..((.....))..)))...((((.((..((....))..))))))(((.......--.)))))))))------------------- ( -25.80, z-score = 0.28, R) >droPer1.super_0 5699919 90 - 11822988 AUCUCGAGCUUCAGGAAGCUGCUGAAUCCCUAUCAGGUUUUCGAUCUGGACAAUGGAGGUCCUCUGCCAGGGUAAAUCCA--AACCAGCAGC------------------- .((..(.....)..)).(((((((((..((.....))..)))...((((.((..((....))..))))))(((.......--.)))))))))------------------- ( -25.50, z-score = -0.09, R) >droWil1.scaffold_181089 11282715 86 - 12369635 AUAUCGAGCUUUAGGAAACUGCUGAAUCCGUAUCAAGUUUUUGACUUGGACAACGGUGGACCAUUGCCGGGGUAAAGCCU--AGACAA----------------------- ...((..(((((((....)).....((((((.(((((((...)))))))))..(((..(....)..))))))))))))..--.))...----------------------- ( -21.90, z-score = -0.71, R) >droVir3.scaffold_13047 18536574 90 + 19223366 AUAUCGAGCUUUAGGAAACUACUGAAUCCGUAUCAGGUCUUUGACCUGGAUAACGGAGGUCCGCUGCCAGGGUGAGUUAA--CCUGAAAACC------------------- .(((.((....(((....))).....)).)))(((((((((...(((((.((.(((....))).)))))))..)))...)--))))).....------------------- ( -27.20, z-score = -1.65, R) >droMoj3.scaffold_6540 7080443 104 + 34148556 AUAUCGAGCUUUCGAAAACUACUUAACCCGUACCAGGUCUUUGACUUGGACAACGGUGGUCCGUUACCCGGGUAAGCGAC--AAUUAAAAUGGAAUGCCUAAGAAC----- ...(((......)))......(((((((((..(((((((...)))))))..(((((....)))))...))))).......--.........((....))))))...----- ( -25.80, z-score = -0.76, R) >droGri2.scaffold_15074 1578845 109 - 7742996 AUAUCGAGCUUUAGAAAACUACUGAACCCGUAUCAGGUCUUCGACUUGGACAACGGAGGACCUCUGCCAGGGUAAUUGUC--AGCGACAAAAAACCAAUCUGCCUGCAUUU .......((..((((..........((((((.(((((((...)))))))))..(((((...)))))...))))..(((((--...)))))........))))...)).... ( -23.70, z-score = 0.07, R) >anoGam1.chr2R 53688429 94 + 62725911 AUUAGUAGUUUUCGUAAAUUAUUAAAUCCAUAUCAAGUGUUUGAUUUGGAUAACGGUGGACCACUGCCUGGGUAUGUUCCUGGGAUUUAUGUUC----------------- .((((((((((....))))))))))(((((.(((((....))))).)))))..(((((...)))))((..((......))..))..........----------------- ( -24.90, z-score = -2.05, R) >apiMel3.Group4 5360165 104 - 10796202 AUCUCCAGCUUUCGAAAACUUUUGAAUCCUUAUCAGGUGUUCGAUCUCGACAAUGGAGGUCCGCUUCCUGGGUUUGUAAAAAAAUUAUAAUACAUCGUAUUUCU------- (((((((....((((......(((((((((....))).))))))..))))...)))))))(((.....)))((((((((.....)))))).))...........------- ( -18.50, z-score = -0.06, R) >triCas2.ChLG2 32668 71 + 12900155 AUAUCCAGUUUUCGAAAGCUUCUGAACCCGUACCAAGUCUUCGAUUUAGACAACGGCGGUCCUUUGCCAGG---------------------------------------- ....((.(((((((((..(((.............)))..)))))...))))...(((((....))))).))---------------------------------------- ( -14.22, z-score = -0.43, R) >consensus AUCUCCAGCUUCAGGAAACUGCUGAAUCCCUAUCAGGUAUUCGAUCUGGACAAUGGCGGUCCUUUGCCAGGGUAAGUUCA__ACUUAAAAAC___________________ .........................((((...((((((.....)))))).....(((((....))))).))))...................................... (-13.75 = -12.80 + -0.95)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:39:21 2011