
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 19,802,858 – 19,802,948 |
| Length | 90 |
| Max. P | 0.551239 |

| Location | 19,802,858 – 19,802,948 |
|---|---|
| Length | 90 |
| Sequences | 15 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 78.71 |
| Shannon entropy | 0.39864 |
| G+C content | 0.54130 |
| Mean single sequence MFE | -36.19 |
| Consensus MFE | -13.09 |
| Energy contribution | -13.20 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.56 |
| Mean z-score | -2.48 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.551239 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
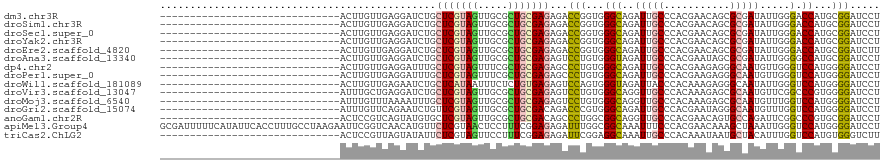
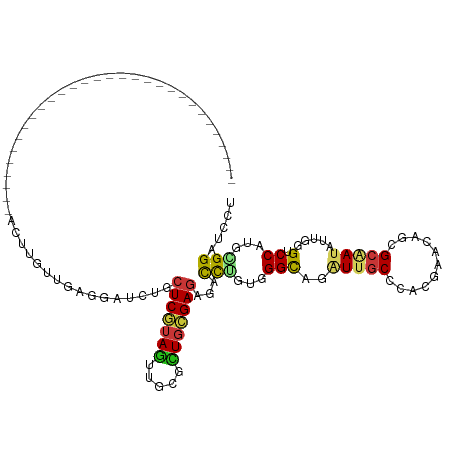
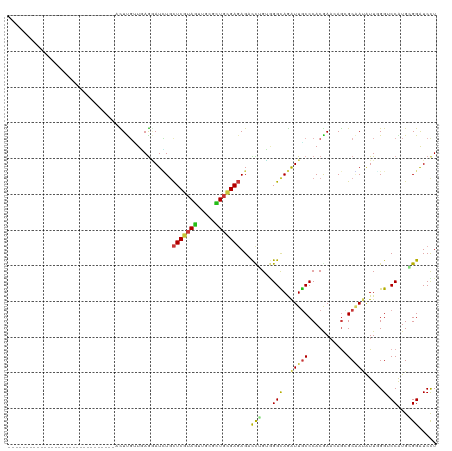
>dm3.chr3R 19802858 90 - 27905053 ------------------------------ACUUGUUGAGGAUCUGCUCGUAGUUGCGCUGCGAGAGACCGGUGGGCAGAUUGCCCACGAACAGCGCGAUAUUGGGACCAUGCGGAUCCU ------------------------------........(((((((((.....(((((((((((......))((((((.....))))))...)))))))))..((....)).))))))))) ( -40.90, z-score = -3.76, R) >droSim1.chr3R 19627091 90 - 27517382 ------------------------------ACUUGUUGAGGAUCUGCUCGUAGUUGCGCUGCGAGAGACCGGUGGGCAGAUUGCCCACGAACAGCGCGAUAUUGGGACCAUGCGGAUCCU ------------------------------........(((((((((.....(((((((((((......))((((((.....))))))...)))))))))..((....)).))))))))) ( -40.90, z-score = -3.76, R) >droSec1.super_0 20144373 90 - 21120651 ------------------------------ACUUGUUGAGGAUCUGCUCGUAGUUGCGCUGCGAGAGACCGGUGGGCAGAUUGCCCACGAACAGCGCGAUAUUGGGACCAUGCGGAUCCU ------------------------------........(((((((((.....(((((((((((......))((((((.....))))))...)))))))))..((....)).))))))))) ( -40.90, z-score = -3.76, R) >droYak2.chr3R 2132944 90 + 28832112 ------------------------------ACUUGUUGAGGAUCUGCUCGUAGUUGCGCUGCGAGAGACCGGUGGGCAGAUUGCCCACGAACAGCGCGAUAUUGGGACCAUGCGGAUCCU ------------------------------........(((((((((.....(((((((((((......))((((((.....))))))...)))))))))..((....)).))))))))) ( -40.90, z-score = -3.76, R) >droEre2.scaffold_4820 2215579 90 + 10470090 ------------------------------ACUUGUUGAGGAUCUGCUCGUAGUUGCGCUGCGAGAGACCGGUGGGCAGAUUGCCCACGAACAGCGCGAUAUUGGGACCAUGCGGAUCUU ------------------------------........(((((((((.....(((((((((((......))((((((.....))))))...)))))))))..((....)).))))))))) ( -38.20, z-score = -3.11, R) >droAna3.scaffold_13340 15941006 90 + 23697760 ------------------------------ACUUGUUGAGGAUCUGCUCGUAGUUGCGCUGCGAGAGUCCUGUGGGUAGAUUGCCCACGAAUAGCGCGAUAUUGGGGCCAUGCGGAUCCU ------------------------------........(((((((((((((((.....))))))(.(((((((((((.....)))))).((((......))))))))))..))))))))) ( -38.30, z-score = -2.87, R) >dp4.chr2 14308723 90 - 30794189 ------------------------------ACUUGUUGAGGAUUUGCUCGUAGUUUCGCUGCGAGAGCCCUGUGGGCAGAUUGCCCACGAAGAGGGCAAUGUUGGGUCCAUGGGGAUCCU ------------------------------........(((((...(((((((.....)))))))..(((((((((((.(((((((.......))))))).)...))))))))))))))) ( -41.50, z-score = -3.61, R) >droPer1.super_0 5699508 90 + 11822988 ------------------------------ACUUGUUGAGGAUUUGCUCGUAGUUUCGCUGCGAGAGCCCUGUGGGCAGAUUGCCCACGAAGAGGGCAAUGUUGGGUCCAUGGGGAUCCU ------------------------------........(((((...(((((((.....)))))))..(((((((((((.(((((((.......))))))).)...))))))))))))))) ( -41.50, z-score = -3.61, R) >droWil1.scaffold_181089 11282314 90 + 12369635 ------------------------------ACUUGUUGAGAAUCUGCUCAUAAUUUCUCUGUGAGAGUCCAGUGGGUAGAUUACCCACAAAGAGGGCAAUAUUGGGUCCAUGGGGAUCCU ------------------------------...(((((..(((((((((((...(((((...)))))....))))))))))).(((.......))))))))..((((((....)))))). ( -30.50, z-score = -1.62, R) >droVir3.scaffold_13047 18536164 90 - 19223366 ------------------------------AUUUGCUGAGGAUCUGCUCGUAGUUGCGCUGCGAGAGUCCUGUGGGCAGGUUGCCCACAAAGAGCGCAAUGUUCGGCCCGUGGGGAUCCU ------------------------------........(((((((((.((..((((((((((....))..(((((((.....)))))))...))))))))...))))(....)))))))) ( -38.70, z-score = -2.02, R) >droMoj3.scaffold_6540 7080037 90 - 34148556 ------------------------------AUUUGUUUAAAAUUUGCUCGUAGUUGCGCUGCGAGAGUCCUGUGGGCAGGUUGCCCACAAAGAGCGCAAUGUUUGGUCCAUGGGGAUCCU ------------------------------......................((((((((((....))..(((((((.....)))))))...))))))))....(((((....))))).. ( -31.30, z-score = -1.66, R) >droGri2.scaffold_15074 1578361 90 + 7742996 ------------------------------AUUUGUUCAGAAUCUGUUCGUAGUUGCGCUGCGACAGACCCGUGGGCAGAUUGCCCACGAAUAGGGCAAUGUUUGGUCCAUGGGGAUCCU ------------------------------............(((((.(((((.....))))))))))((((((((((.(((((((.......))))))).)...)))))))))...... ( -39.30, z-score = -3.81, R) >anoGam1.chr2R 53688009 90 - 62725911 ------------------------------ACUCCGUCAGUAUGUGCUCGUAGUUGCGCUGCGACAGCCCUGGCGGCAGGUUGCCCACGAACAGUGCCAGAUUCGGCCCGUGCGGAUCCU ------------------------------...(((((((...((..((((((.....))))))..)).))))))).(((...((((((....(.(((......)))))))).))..))) ( -31.70, z-score = 0.31, R) >apiMel3.Group4 5359677 120 + 10796202 GCGAUUUUUCAUAUUCACCUUUGCCUAAGAAUUCGGUCAACAUGUUCUCGUAACUCCUUUCGGAGAGAUUUGGCGGCAAAUUUCCCACGAACAAAGCUAAAUUGGGUCCAUGGGGAUCCU ((((................))))..........((((..((((..(((....((((....)))).((((((((.....................)))))))))))..))))..)))).. ( -26.99, z-score = 0.42, R) >triCas2.ChLG2 31691 90 - 12900155 ------------------------------ACUCCGUUAGUAUAUUCUCGUAGUUCCUUUCGGAGAGAUUCGGAGGCAAAUUGCCCACAAAUAAUGCUACAUUUGGUCCAUGUGGGUCUU ------------------------------.(((((.........((((.....(((....)))))))..))))).......(((((((......((((....))))...)))))))... ( -21.20, z-score = -0.55, R) >consensus ______________________________ACUUGUUGAGGAUCUGCUCGUAGUUGCGCUGCGAGAGACCUGUGGGCAGAUUGCCCACGAACAGCGCAAUAUUGGGUCCAUGCGGAUCCU ..............................................(((((((.....)))))))...(((...(((..(((((...........))))).....).))...)))..... (-13.09 = -13.20 + 0.11)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:39:20 2011