| Sequence ID | dm3.chr3R |
|---|---|
| Location | 19,736,123 – 19,736,191 |
| Length | 68 |
| Max. P | 0.999890 |

| Location | 19,736,123 – 19,736,191 |
|---|---|
| Length | 68 |
| Sequences | 11 |
| Columns | 78 |
| Reading direction | reverse |
| Mean pairwise identity | 61.69 |
| Shannon entropy | 0.78217 |
| G+C content | 0.41322 |
| Mean single sequence MFE | -21.21 |
| Consensus MFE | -15.60 |
| Energy contribution | -15.35 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.54 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 4.73 |
| SVM RNA-class probability | 0.999890 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
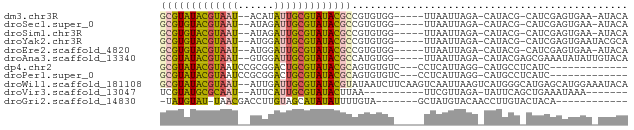
>dm3.chr3R 19736123 68 - 27905053 GCGUAUACGUAAU--ACAUAUUGCGUAUACGCCGUGUGG-----UUAAUUAGA-CAUACG-CAUCGAGUGAA-AUACA (((((((((((((--....))))))))))))).((((((-----((.....))-).))))-)..........-..... ( -23.30, z-score = -2.91, R) >droSec1.super_0 20077867 68 - 21120651 GCGUGUACGUAAU--AUAGAUUGCGUAUACGCCGUGUGG-----UUAAUUAGA-CAUACG-CAUCGAGUGAA-AUACA (((((((((((((--....))))))))))))).((((((-----((.....))-).))))-)..........-..... ( -23.40, z-score = -2.79, R) >droSim1.chr3R 19556912 68 - 27517382 GCGUGUACGUAAU--AUAGAUUGCGUAUACGCCGUGUGG-----UUAAUUAGA-CAUACG-CAUCGAGUGAA-AUACA (((((((((((((--....))))))))))))).((((((-----((.....))-).))))-)..........-..... ( -23.40, z-score = -2.79, R) >droYak2.chr3R 20629094 69 - 28832112 GCGUGUACGUAAU--AUGGAUUGCGUAUACGCCGUGUGG-----UUAAUUAGA-CAUACG-CAUCGAGUGAAUACGCA (((((((((((((--....))))))))))))).((((((-----((.....))-).))))-).....(((....))). ( -24.70, z-score = -2.04, R) >droEre2.scaffold_4820 2147994 68 + 10470090 GCGUGUACGUAAU--AUGGAUUGCGUAUACGCCGUGUGG-----UUAAUUAGA-CAUACG-CAUCGAGUGAA-AUACA (((((((((((((--....))))))))))))).((((((-----((.....))-).))))-)..........-..... ( -23.40, z-score = -2.41, R) >droAna3.scaffold_13340 15875439 70 + 23697760 GCGUAUACGUAAU--GUGGAUUGCGUAUACGCCAUGUGG-----UUAAUUAGA-CAUACGAGCGAAAUAUAUUGUACA (((((((((((((--....)))))))))))))...((((-----((.....))-).)))(.((((......)))).). ( -20.30, z-score = -1.68, R) >dp4.chr2 16881763 61 + 30794189 GCGUAUACGUAAUCCGCGGACUGCGUAUACGCAGUGUGUC---CCUCAUUAGG-CAUGCCUCAUC------------- (((((((((((.((....)).))))))))))).(((((((---........))-)))))......------------- ( -25.60, z-score = -4.12, R) >droPer1.super_0 6532640 61 - 11822988 GCGUAUACGUAAUCCGCGGACUGCGUAUACGCAGUGUGUC---CCUCAUUAGG-CAUGCCUCAUC------------- (((((((((((.((....)).))))))))))).(((((((---........))-)))))......------------- ( -25.60, z-score = -4.12, R) >droWil1.scaffold_181108 3959907 76 + 4707319 GCGUAUACGUAAU--AUUGAUUGCGUAUACGUAUAAUCUUCAAGUCAAUUAAGUCAUGGGCAUGAGCAUGGAAAUACA (((((((((((((--....))))))))))))).....................(((((........)))))....... ( -19.20, z-score = -2.14, R) >droVir3.scaffold_13047 13973409 58 - 19223366 UCGUAUGCGCAAU--AUUCAUUGCGUAUACUUAA----------UUCGUUAGA-UAUUCAGCUGAAAUAAA------- ..(((((((((((--....)))))))))))(((.----------((((((.(.-....)))).))).))).------- ( -14.90, z-score = -2.97, R) >droGri2.scaffold_14830 283909 57 + 6267026 -UAUGUAU-UAACGACCUUGUAGCAUAUAUUUUGUA-------GCUAUGUACAACCUUGUACUACA------------ -..((((.-..........((((((((.....))).-------)))))(((((....)))))))))------------ ( -9.50, z-score = -0.69, R) >consensus GCGUAUACGUAAU__AUGGAUUGCGUAUACGCCGUGUGG_____UUAAUUAGA_CAUACG_CAUCGAGUGAA_AUACA (((((((((((((......))))))))))))).............................................. (-15.60 = -15.35 + -0.24)

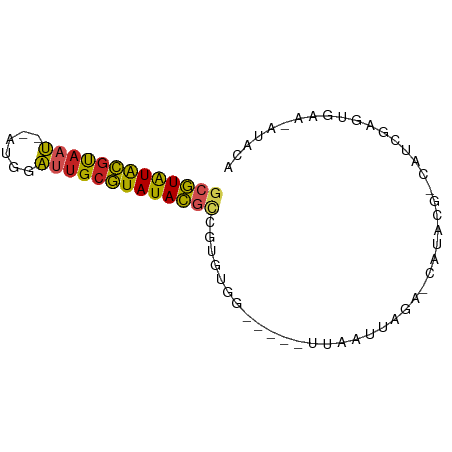
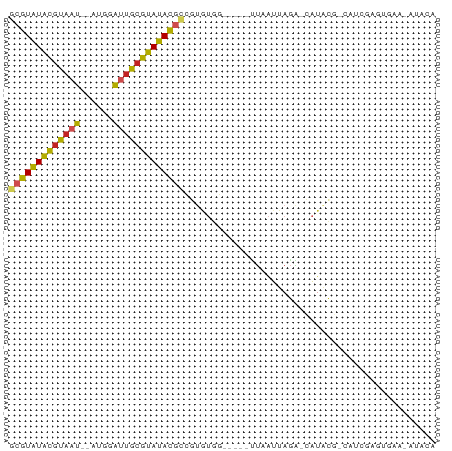
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:39:07 2011