
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 19,725,163 – 19,725,263 |
| Length | 100 |
| Max. P | 0.836080 |

| Location | 19,725,163 – 19,725,263 |
|---|---|
| Length | 100 |
| Sequences | 12 |
| Columns | 100 |
| Reading direction | forward |
| Mean pairwise identity | 68.59 |
| Shannon entropy | 0.67188 |
| G+C content | 0.55343 |
| Mean single sequence MFE | -35.32 |
| Consensus MFE | -13.78 |
| Energy contribution | -13.52 |
| Covariance contribution | -0.26 |
| Combinations/Pair | 1.56 |
| Mean z-score | -1.44 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.85 |
| SVM RNA-class probability | 0.836080 |
| Prediction | RNA |
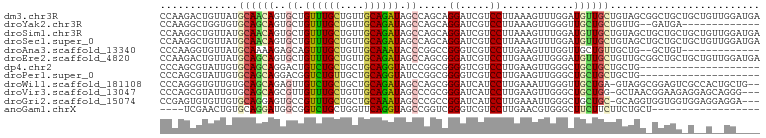
Download alignment: ClustalW | MAF
>dm3.chr3R 19725163 100 + 27905053 CCAAGACUGUUAUGCAACAGUGCUGUUUGCUGUUGCAGAUAGCCAGCAGGAUCGUCCUUAAAGUUUGGAUGUUGCUGUAGCGGCUGCUGCUGUUGGAUGA ((((.((((((....))))))(((((((((....)))))))))((((((....((((.........))))(((((....)))))..)))))))))).... ( -38.10, z-score = -2.36, R) >droYak2.chr3R 20617433 85 + 28832112 CCAAGGCUGGUGUGCAGCAGUGCUGUUUGCUGUUGCAGAUAGCCAGCAGGAUCGUCCUUAAAGUUGGGUUGCUGCUGUUG--GAUGA------------- (((((((((.(.(((((((((.......))))))))).))))))(((((.(...(((........))).).))))).)))--)....------------- ( -34.20, z-score = -2.02, R) >droSim1.chr3R 19546039 100 + 27517382 CCAAGGCUGUUAUGCAACAGUGCUGUUUGCUGUUGCAGAUAGCCAGCAGGAUCGUCCUUAAAGUUUGGAUGUUGCUGUAGCUGCUGCUGCUGUUGGAUGA (((((((((((.(((((((((.......))))))))))))))))(((((...(((((.........))))))))))(((((....)))))..)))).... ( -42.00, z-score = -3.31, R) >droSec1.super_0 20067032 100 + 21120651 CCAAGGCUGUUAUGCAACAGUGCUGUUUGCUGUUGCAGAUAGCCAGCAGGAUCGUCCUUAAAGUUUGGAUGUUGCUGUAGCUGCUGCUGCUGUUGGAUGA (((((((((((.(((((((((.......))))))))))))))))(((((...(((((.........))))))))))(((((....)))))..)))).... ( -42.00, z-score = -3.31, R) >droAna3.scaffold_13340 15864328 85 - 23697760 CCCAAGGUGUUAUGCAAAAGAGCAGUUUGCUGUUGCAAAUACCCGGCCGGGUCGUCCUUGAAGUUUGGUUGCUGUUGCUG--GCUGU------------- .....((((((.(((((...(((.....))).)))))))))))(((((((.(((....)))(((......)))....)))--)))).------------- ( -25.90, z-score = -0.43, R) >droEre2.scaffold_4820 2136774 100 - 10470090 CCAAGACUGUUAUGCAGCAGUGCUGUUUGCUGUUGCAGAUAGCCAGCGGGAUCGUCCUUGAAGUUGGGAUGUUGCUGUUGCGGCUGCUGCUGUUGGAUGA ((((.(.......(((((((((((((((((....)))))))))((((((.(.((((((.......)))))).).))))))..))))))))).)))).... ( -43.81, z-score = -3.50, R) >dp4.chr2 16869330 80 - 30794189 CCCAGCGUAUUGUGCAGCAGGACUGUCUGCUGCUGCAGGUAUCCGGCGGGGUCGUCCUUGAAGUUGGGCUGCUGCUGCUG-------------------- ..((((.......((((((((....)))))))).((((.((((((((.(((....)))....)))))).)))))).))))-------------------- ( -31.30, z-score = -0.02, R) >droPer1.super_0 6519440 80 + 11822988 CCCAGCGUAUUGUGCAGCAGGACGGUCUGUUGCUGCAGGUAUCCGGCGGGGUCGUCCUUGAAGUUGGGCUGCUGCUGCUG-------------------- ..(((((((..((.((((((((((..((..(((((........)))))))..))))))....)))).))...))).))))-------------------- ( -30.50, z-score = 0.02, R) >droWil1.scaffold_181108 3948571 97 - 4707319 CCCAGGGUGUUGUGCAGCAGAGUUGUCUGCUGCUGCAGAUAGCCAGCGGGAUCAUCCUUGAAAUUGGGUUGCUGA-GUAGGCGGAGUCGCCACUGCUG-- ((((((((.((((((((((((....)))))))).))))...)))...(((.....))).....)))))..((..(-((.((((....)))))))))..-- ( -36.90, z-score = -0.61, R) >droVir3.scaffold_13047 13962673 96 + 19223366 CCCAGCGUAUUGUGCAGCAGCGUUGUUUGCUGUUGCAGAUAGCCCGCGGGAUCAUCCUUGAAGUUGGGCUGCUGG-GCUAACGGAAGAGGAGCAGGG--- .(((((....(.((((((((((.....)))))))))).).((((((((((.....)))....).)))))))))))-(((..(....)...)))....--- ( -40.80, z-score = -2.10, R) >droGri2.scaffold_15074 7657169 96 - 7742996 CCGAGUGUGUUGUGCAGGAGUGCCGUUUGCUGCUGCAAAUAGCCCGCCGGAUCAUCCUUGAAAUUGGGCUGCUGC-GCAGGUGGUGGUGGAGGAGGA--- (((.(((.(((((((((.(((.......))).)))))..)))).)))))).((.(((((.(...(.(.((((...-)))).).)...).))))).))--- ( -32.40, z-score = 0.44, R) >anoGam1.chrX 9484030 78 - 22145176 ----UCGAACUGUGCAGGAUGGCGGUCUGCUGGUUCAGGUAGCCGGUCGGGUCGUCCUUGAACGUGGGCUUCUUCUUCUGCU------------------ ----.........((((((....(..(((((((((.....)))))).)))..)((((........))))......)))))).------------------ ( -25.90, z-score = -0.11, R) >consensus CCAAGGGUGUUGUGCAGCAGUGCUGUUUGCUGUUGCAGAUAGCCAGCGGGAUCGUCCUUGAAGUUGGGCUGCUGCUGCUGC_GCUGC_G__G__GG____ .............((((((..((.((((((....)))))).)).....((.....))............))))))......................... (-13.78 = -13.52 + -0.26)


Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:39:05 2011