
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 19,404,377 – 19,404,483 |
| Length | 106 |
| Max. P | 0.579182 |

| Location | 19,404,377 – 19,404,483 |
|---|---|
| Length | 106 |
| Sequences | 14 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 78.00 |
| Shannon entropy | 0.48226 |
| G+C content | 0.62975 |
| Mean single sequence MFE | -43.23 |
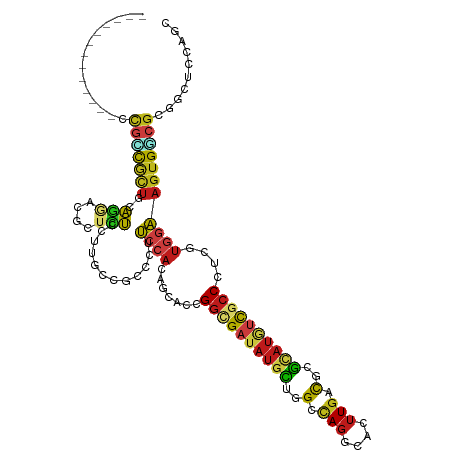
| Consensus MFE | -18.90 |
| Energy contribution | -19.49 |
| Covariance contribution | 0.59 |
| Combinations/Pair | 1.71 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.44 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.579182 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
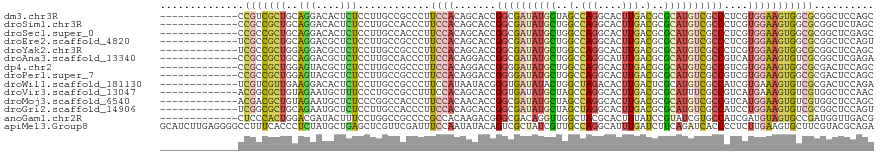
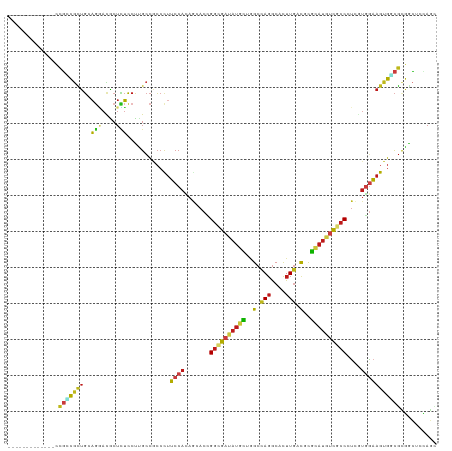
>dm3.chr3R 19404377 106 - 27905053 -------------CCGUCGCUGCAGGACACUCUCCUUGCCGCCCUUCCACAGCACCGGCGAUAUGCUAGCCAGGCACUUGACGCGCAUGUCGCCCUCGUGGAAGUGGCGCGGCUCCAGC -------------..(((((.(((((........))))).((((((((((((....((((((((((..(.(((....))).)..)))))))))))).))))))).))))))))...... ( -48.10, z-score = -2.60, R) >droSim1.chr3R 19226427 106 - 27517382 -------------CCGCCGCUGCAGGACACUCUCCUUGCCACCCUUCCACAGCACCGGCGAUAUGCUGGCCAGGCACUUGACGCGCAUGUCGCCCUCGUGGAAGUGGCGCGGCUCUAGC -------------..(((((...((((.....)))).(((((..((((((((....((((((((((((..(((....))).)).)))))))))))).))))))))))))))))...... ( -48.10, z-score = -2.51, R) >droSec1.super_0 19763473 106 - 21120651 -------------CCGCCGCUGCAGGACACUCUCCUUGCCACCCUUCCACAGCACCGGCGAUAUGCUGGCCAGGCACUUGACGCGCAUGUCGCCCUCGUGGAAGUGGCGCGGCUCGAGC -------------..(((((...((((.....)))).(((((..((((((((....((((((((((((..(((....))).)).)))))))))))).))))))))))))))))...... ( -48.10, z-score = -2.18, R) >droEre2.scaffold_4820 1830307 106 + 10470090 -------------UCGCCGCUGCAGGACGCUCUCCUUGCCGCCCUUCCACAGCACCGGCGAUAUGCUGGCCAGGCACUUGACGCGCAUGUCGCCCUCGUGGAAGUGGCGCGGCUCCAGU -------------..(((((.(((((........))))).((((((((((((....((((((((((((..(((....))).)).)))))))))))).))))))).))))))))...... ( -51.60, z-score = -2.50, R) >droYak2.chr3R 20290255 106 - 28832112 -------------UCGCCGCUGGAGGACGCUCUCCUUGCCGCCCUUCCACAGCACCGGCGAUAUGCUGGCCAGGCACUUGACGCGCAUGUCGCCCUCGUGGAAGUGGCGCGGCUCCAGC -------------..(((((.(((((....))))).....((((((((((((....((((((((((((..(((....))).)).)))))))))))).))))))).))))))))...... ( -52.10, z-score = -2.38, R) >droAna3.scaffold_13340 3553913 106 + 23697760 -------------CCGCCGCUGCAGGACGCUCUCCUUGCCACCCUUCCACAGGACCGGCGAUAUGCUGGCCAGGCAUUUGACGCGCAUGUCGCCGUCAUGGAAGUGUCGCGGCUCGAGA -------------..(((((.(((((........))))).((.((((((...(((.((((((((((((..(((....))).)).))))))))))))).)))))).)).)))))...... ( -45.80, z-score = -1.60, R) >dp4.chr2 2891197 106 - 30794189 -------------CCGCCGCUGGAGUACGCUCUCCUUGCCGCCCUUCCACAGGACCGGGGAUAUGCUGGCCAGGCACUUGACGCGCAUGUCGCCGUCGUGGAAGUGGCGCGACUCCAGC -------------.....((((((((.(((.(.....)..((((((((((..(((.((.(((((((((..(((....))).)).))))))).)))))))))))).)))))))))))))) ( -51.40, z-score = -2.36, R) >droPer1.super_7 2984739 106 + 4445127 -------------CCGCCGCUGGAGUACGCUCUCCUUGCCGCCCUUCCACAGGACCGGGGAUAUGCUGGCCAGGCACUUGACGCGCAUGUCGCCGUCGUGGAAGUGGCGCGACUCCAGC -------------.....((((((((.(((.(.....)..((((((((((..(((.((.(((((((((..(((....))).)).))))))).)))))))))))).)))))))))))))) ( -51.40, z-score = -2.36, R) >droWil1.scaffold_181130 5459253 106 - 16660200 -------------UCGUCGUUGAAGGACACUCUCCUUGCCGCCCUUCCAUAAUACGGGUGAUAUACUGGCUAGACACUUGACUCGCAUGUCGCCAUCGUGAAAGUGUCGCGACUCCAGA -------------..(((((..(((((.....)))))..(((((...........)))))............(((((((...(((((((....))).))))))))))))))))...... ( -29.30, z-score = -0.43, R) >droVir3.scaffold_13047 9019695 106 - 19223366 -------------ACGGCGCUGUAGAAUGCUUUCCCUGCCGCCUUUCCACAGCACCGGUGAUAUGCUAGCCAGGCACUUGACGCGCAUUUCGCCGUCAUGAAAGUGUCGUGGCUCCAAC -------------.(((.(((((.(((.((..........))..))).))))).)))..........((((((((((((((((.((.....)))))))....)))))).)))))..... ( -38.80, z-score = -2.11, R) >droMoj3.scaffold_6540 2514133 106 + 34148556 -------------ACGACGCUGUAGAAUGCUCUCCCGGCCACCCUUCCACAACACCGGCGAUAUGCUAGCCAGGCACUUGACGCGCAUGUCGCCGUCAUGGAAGUGUCGUGGCUCCAGC -------------.....((((.(((....)))...(((((((((((((......(((((((((((..(.(((....))).)..)))))))))))...)))))).)..)))))).)))) ( -38.70, z-score = -1.37, R) >droGri2.scaffold_14906 9945195 106 + 14172833 -------------UCGGCGCUGCAGAAUGCUCUCCUUGCCACCCUUCCACAGCACCGGCGAUAUGCUAGCUAGGCACUUGACGCGCAUGUCGCCAUCCUGGAAGUGUCGCGGCUCCAGU -------------..((.(((((.....((.......)).((.((((((.......((((((((((..((............))))))))))))....)))))).)).))))).))... ( -36.70, z-score = -0.22, R) >anoGam1.chr2R 53129466 106 + 62725911 -------------CUCCCACUGGACGAUACUUUCCUGGCCGCCCCGCCACAAGACGGGCGACAGGUUGGCUACGCACUUUAUCCGUAUCGUGCCAUCGAUGUAGUGCCGAUGGUUGACG -------------..........(((((((...((((..(((((..(.....)..))))).))))..((.((.......)).)))))))))(((((((.........)))))))..... ( -32.00, z-score = 0.10, R) >apiMel3.Group8 5808374 119 - 11452794 GCAUCUUGAGGGGCCUUUCACCCUCUAUGCUGAGCUCGUUCGAUUUCCAAUAUACAGUCGCUAUCGUUGCCAGGCAUUUGAUCUUCAGAUCACCCCUCUUGAAGUGCUUCGUACGCAGA ((.((..((((((.............(((((..((.((..(((((..........)))))....))..))..))))).(((((....)))))))))))..)).((((...))))))... ( -33.10, z-score = -1.43, R) >consensus _____________CCGCCGCUGCAGGACGCUCUCCUUGCCGCCCUUCCACAGCACCGGCGAUAUGCUGGCCAGGCACUUGACGCGCAUGUCGCCCUCGUGGAAGUGGCGCGGCUCCAGC ..............(((((((..(((....)))............((((.......((((((((((..(.(((....))).)..))))))))))....))))))))))).......... (-18.90 = -19.49 + 0.59)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:38:28 2011