
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 19,259,740 – 19,259,872 |
| Length | 132 |
| Max. P | 0.762477 |

| Location | 19,259,740 – 19,259,815 |
|---|---|
| Length | 75 |
| Sequences | 5 |
| Columns | 77 |
| Reading direction | forward |
| Mean pairwise identity | 95.31 |
| Shannon entropy | 0.08147 |
| G+C content | 0.34212 |
| Mean single sequence MFE | -11.23 |
| Consensus MFE | -11.32 |
| Energy contribution | -11.16 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.07 |
| Mean z-score | -0.82 |
| Structure conservation index | 1.01 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.688604 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
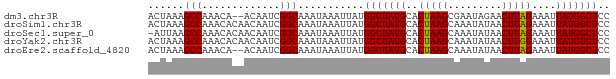
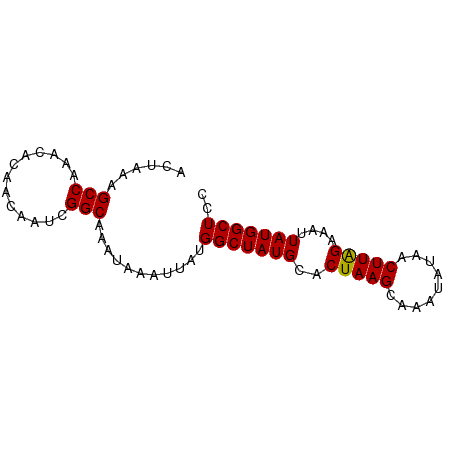
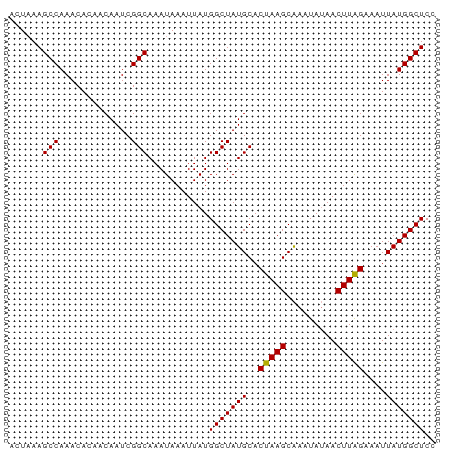
>dm3.chr3R 19259740 75 + 27905053 ACUAAAGCCAAACA--ACAAUCGGCAAAUAAAUUAUGGCUAUGCACUAAGCGAAUAGAACUUAGAAAUUAUGGCUCC ......(((.....--......)))...........(((((((..(((((.........)))))....))))))).. ( -11.40, z-score = -0.77, R) >droSim1.chr3R 19077748 77 + 27517382 ACUAAAGCCAAACACAACAAUCGGCAAAUAAAUUAUGGCUAUGCACUAAGCAAAUAUAACUUAGAAAUUAUGGCUCC ......(((.............)))...........(((((((..(((((.........)))))....))))))).. ( -11.22, z-score = -1.02, R) >droSec1.super_0 19622634 76 + 21120651 -AUUAAGCCAAACACAACAAUCGGCAAAUAAAUUAUGGCUAUGCACUAAGCAAAUAUAACUUAGAAAUUAUGGCUCC -.....(((.............)))...........(((((((..(((((.........)))))....))))))).. ( -11.22, z-score = -0.85, R) >droYak2.chr3R 20141582 77 + 28832112 ACUAAAGCCAAACACAACAAUCGGCAAAUAAAUUAUGGCUAUGCACUAAGCAAAUAUAACUUGGAAAUUAUGGCUCC ......(((.............)))...........(((((((..(((((.........)))))....))))))).. ( -10.92, z-score = -0.49, R) >droEre2.scaffold_4820 1680108 75 - 10470090 ACUAAAGCCAAACA--ACAAUCGGCAAAUAAAUUAUGGCUAUGCACUAAGCAAAUAUAACUUAGAAAUUAUGGCUCC ......(((.....--......)))...........(((((((..(((((.........)))))....))))))).. ( -11.40, z-score = -0.97, R) >consensus ACUAAAGCCAAACACAACAAUCGGCAAAUAAAUUAUGGCUAUGCACUAAGCAAAUAUAACUUAGAAAUUAUGGCUCC ......(((.............)))...........(((((((..(((((.........)))))....))))))).. (-11.32 = -11.16 + -0.16)
| Location | 19,259,815 – 19,259,872 |
|---|---|
| Length | 57 |
| Sequences | 6 |
| Columns | 57 |
| Reading direction | reverse |
| Mean pairwise identity | 78.36 |
| Shannon entropy | 0.41969 |
| G+C content | 0.39059 |
| Mean single sequence MFE | -10.32 |
| Consensus MFE | -6.26 |
| Energy contribution | -7.78 |
| Covariance contribution | 1.53 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.54 |
| Structure conservation index | 0.61 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.61 |
| SVM RNA-class probability | 0.762477 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 19259815 57 - 27905053 GUAUAACAAAUGUGCGGCAAAUGAGCUGUCAUAGGUAAAUAAGGAAGACGAGAAGCU .....((...((((((((......)))).)))).))..................... ( -8.60, z-score = -0.60, R) >droEre2.scaffold_4820 1680183 57 + 10470090 GUAUAACAAAUCUGCGGCAAAUGAGCUGUCGUAGGUAAAUAAGGAAGACGAGAAACC .........((((((((((.......))))))))))..................... ( -12.10, z-score = -2.09, R) >droYak2.chr3R 20141659 57 - 28832112 GUAUAACAAAUCUGCGGCAAAUGAGCUGUCGUAGGUAAAUAAGGAAGACGAGAAACC .........((((((((((.......))))))))))..................... ( -12.10, z-score = -2.09, R) >droSec1.super_0 19622710 57 - 21120651 UUAUAACAAAUCUGCGGCAAAUGAGCUGUCGUAGGUAAAUAAGGAAGACGAGAAACC .........((((((((((.......))))))))))..................... ( -12.10, z-score = -2.16, R) >droSim1.chr3R 19077825 57 - 27517382 UUAUAACAAAUCUGCGGCAAAUGAGCUGUCGUAGGUAAAUAAGGAAGACGAGAAACC .........((((((((((.......))))))))))..................... ( -12.10, z-score = -2.16, R) >droWil1.scaffold_181130 9299628 55 - 16660200 UAAAGACUGAACCUCAAAAAGAACACGGUCA-AAGUAAUUUAAAUGGAAUAGUAUU- ....(((((....((.....))...))))).-........................- ( -4.90, z-score = -0.12, R) >consensus GUAUAACAAAUCUGCGGCAAAUGAGCUGUCGUAGGUAAAUAAGGAAGACGAGAAACC .........((((((((((.......))))))))))..................... ( -6.26 = -7.78 + 1.53)



Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:37:52 2011