
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 18,873,319 – 18,873,430 |
| Length | 111 |
| Max. P | 0.737482 |

| Location | 18,873,319 – 18,873,429 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 64.88 |
| Shannon entropy | 0.57239 |
| G+C content | 0.42955 |
| Mean single sequence MFE | -28.85 |
| Consensus MFE | -14.07 |
| Energy contribution | -18.07 |
| Covariance contribution | 4.00 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.09 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.29 |
| SVM RNA-class probability | 0.631927 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 18873319 110 - 27905053 GGGCUAACAAAUGGCUGCAAUGAGCGUCUGGCAAAUGAGCCAUUAAUAAGGCUA---GUCAGAUGCACAUCAGA-CAUGGAUGCACUUAGAAAAUGCAGUCGCAUUUCAUGUUA .(((((.....)))))...(((.(((((((((.....((((........)))).---))))))))).)))..((-((((((((((((((.....)).))).))).)))))))). ( -36.00, z-score = -2.48, R) >droYak2.chr3R 19737193 110 - 28832112 GUGCUAACAAAUGGCUGCAAUGAACGCCUGGCAAAUGAGCCAUUAAUAAGGCUA---GUCAGAUACACAUCGGA-CUUGGAUGCACUCGGAAAAUGCAGUAUGCUUUCACGUUA ((((...(((.(((((((.......)).((((......)))).......)))))---(((.(((....))).))-))))...)))).((((((..((.....)))))).))... ( -24.40, z-score = 0.71, R) >droSim1.chr3R 18683953 98 - 27517382 GGGCUAACAAAUGGCUGCAAUGAGCGUCUGGCAAAUGAGCCAUUAAUAAGGCUA---GUCAGAUGCACAUCAGA-CAUGGAUGCACUUAGAAAAUGCAGUCU------------ ............(((((((....(((((((((.....((((........)))).---))))))))).((((...-....))))...........))))))).------------ ( -32.70, z-score = -2.68, R) >droGri2.scaffold_15074 3685347 111 - 7742996 AUGCGAGUAAA--ACUGUAUUGAAAGCUUUACCUAUAAG-CAUUACAGCGUAUACAUAUCUUAUACAAAUAGGAGUGUGAGAGAGAGAGAGAGGGGGAGUUGGUUGCUGUACUA .....((((..--...(((.....(((((..(((....(-(......)).........((((((((........)))))))).........)))..)))))...)))..)))). ( -22.30, z-score = 0.10, R) >consensus GGGCUAACAAAUGGCUGCAAUGAACGCCUGGCAAAUGAGCCAUUAAUAAGGCUA___GUCAGAUACACAUCAGA_CAUGGAUGCACUUAGAAAAUGCAGUCGG_UUUCAUGUUA ............(((((((....(((((((((.....((((........))))....))))))))).((((........))))...........)))))))............. (-14.07 = -18.07 + 4.00)


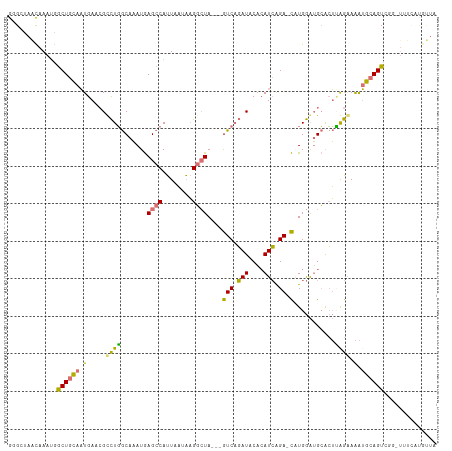
| Location | 18,873,320 – 18,873,430 |
|---|---|
| Length | 110 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 64.58 |
| Shannon entropy | 0.57843 |
| G+C content | 0.43548 |
| Mean single sequence MFE | -29.34 |
| Consensus MFE | -14.07 |
| Energy contribution | -18.07 |
| Covariance contribution | 4.00 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.23 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.737482 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 18873320 110 - 27905053 GGGGCUAACAAAUGGCUGCAAUGAGCGUCUGGCAAAUGAGCCAUUAAUAAGGCUA---GUCAGAUGCACAUCAGA-CAUGGAUGCACUUAGAAAAUGCAGUCGCAUUUCAUGUU (.(((((.....))))).).(((.(((((((((.....((((........)))).---))))))))).)))..((-((((((((((((((.....)).))).))).)))))))) ( -37.40, z-score = -2.88, R) >droYak2.chr3R 19737194 110 - 28832112 CGUGCUAACAAAUGGCUGCAAUGAACGCCUGGCAAAUGAGCCAUUAAUAAGGCUA---GUCAGAUACACAUCGGA-CUUGGAUGCACUCGGAAAAUGCAGUAUGCUUUCACGUU ((((..(.((....((((((.....((....(((....((((........))))(---(((.(((....))).))-))....)))...)).....)))))).)).)..)))).. ( -25.60, z-score = 0.37, R) >droSim1.chr3R 18683953 99 - 27517382 GGGGCUAACAAAUGGCUGCAAUGAGCGUCUGGCAAAUGAGCCAUUAAUAAGGCUA---GUCAGAUGCACAUCAGA-CAUGGAUGCACUUAGAAAAUGCAGUCU----------- .............(((((((....(((((((((.....((((........)))).---))))))))).((((...-....))))...........))))))).----------- ( -32.70, z-score = -2.58, R) >droGri2.scaffold_15074 3685348 111 - 7742996 UAUGCGAGUAAA--ACUGUAUUGAAAGCUUUACCUAUAAG-CAUUACAGCGUAUACAUAUCUUAUACAAAUAGGAGUGUGAGAGAGAGAGAGAGGGGGAGUUGGUUGCUGUACU ..(((.(((((.--...........(((((..(((....(-(......)).........((((((((........)))))))).........)))..)))))..)))))))).. ( -21.64, z-score = 0.17, R) >consensus GGGGCUAACAAAUGGCUGCAAUGAACGCCUGGCAAAUGAGCCAUUAAUAAGGCUA___GUCAGAUACACAUCAGA_CAUGGAUGCACUUAGAAAAUGCAGUCGG_UUUCAUGUU .............(((((((....(((((((((.....((((........))))....))))))))).((((........))))...........)))))))............ (-14.07 = -18.07 + 4.00)


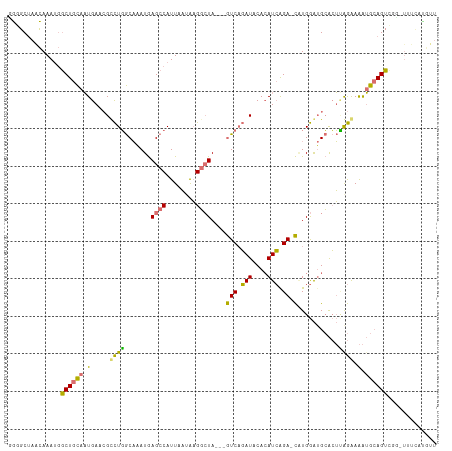
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:37:08 2011