
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 18,845,594 – 18,845,699 |
| Length | 105 |
| Max. P | 0.719585 |

| Location | 18,845,594 – 18,845,699 |
|---|---|
| Length | 105 |
| Sequences | 9 |
| Columns | 107 |
| Reading direction | reverse |
| Mean pairwise identity | 95.53 |
| Shannon entropy | 0.09857 |
| G+C content | 0.38013 |
| Mean single sequence MFE | -24.86 |
| Consensus MFE | -20.51 |
| Energy contribution | -21.03 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.719585 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 18845594 105 - 27905053 -GUCAGGCAAAAAUAAAGUGGCCAGAAG-GAUGGUCCCGUUAAAUGAGCAGUUAACAUGGCCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC -((((((((.((((...(.(((((...(-(......))(((((........))))).))))))(((((..............))))).)))).))))..)))).... ( -23.24, z-score = -1.21, R) >droSim1.chr3R 18659693 105 - 27517382 -GUCAGGCAAAAAUAAAGUGGCCAGAAG-CAUGGCCCCGUUAAAUGAGCAGUUAACAUUGCCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC -((((((((.((((...(.(((((....-..))))).)(((.((((.(((((....))))).))))...)))................)))).))))..)))).... ( -25.50, z-score = -2.38, R) >droSec1.super_0 19209270 105 - 21120651 -GUCAGGCAAAAAUAAAGUGGCCAGAAG-CAUGGCCCCGUUAAAUGAGCAGUUAACAUUGCCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC -((((((((.((((...(.(((((....-..))))).)(((.((((.(((((....))))).))))...)))................)))).))))..)))).... ( -25.50, z-score = -2.38, R) >droYak2.chr3R 19707668 105 - 28832112 -GUCAGGCAAAAAUAAAGUGGCCAGAAG-CAUGGCCCUGUUAAAUGAGCAGUUAACAUUGCCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC -((((((((.((((..((.(((((....-..)))))))(((.((((.(((((....))))).))))...)))................)))).))))..)))).... ( -25.90, z-score = -2.27, R) >droEre2.scaffold_4820 1246259 105 + 10470090 -GUCAGGCUUAAAUAAAGUGGCCAGAAG-CAUGGCCCCGUUAAAUGAGCAGUUAACAUUGCCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC -(((((((.(((((...(.(((((....-..))))).)(((.((((.(((((....))))).))))...)))................))))).)))..)))).... ( -25.30, z-score = -2.14, R) >droAna3.scaffold_13340 11224255 105 + 23697760 -GUCAGGCAAAAAUAAAGUGGCCAGAAG-CAUGGCCCCGUUAAAUGAGCAGUUAACACUGCCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC -((((((((.((((...(.(((((....-..))))).)(((.((((.(((((....))))).))))...)))................)))).))))..)))).... ( -27.80, z-score = -3.08, R) >dp4.chr2 24764004 106 + 30794189 -GUCAGGCAAAAAUAAAGUGGCCAGAAGCCAUGGCCCCGUUAAAUGAGCAGUUAACAUUGGCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC -((((((((.((((...(((((.....)))))((((..(((((........)))))...))))(((((..............))))).)))).))))..)))).... ( -25.54, z-score = -1.85, R) >droPer1.super_19 101081 106 + 1869541 -GUCAGGCAAAAAUAAAGUGGCCAGAAGCCAUGGCCCCGUUAAAUGAGCAGUUAACAUUGGCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC -((((((((.((((...(((((.....)))))((((..(((((........)))))...))))(((((..............))))).)))).))))..)))).... ( -25.54, z-score = -1.85, R) >droWil1.scaffold_181089 11187172 105 + 12369635 GAUGUAGUACAAAUAAAGUGGCCAGAGU-CAUGG-CCCGUAAAAUGAGCAGUUAACAUUGCCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC ..(((........(((((((((((....-..)))-))((((..(((.(((((....))))).))))))).))))))...........(((........))))))... ( -19.40, z-score = -0.73, R) >consensus _GUCAGGCAAAAAUAAAGUGGCCAGAAG_CAUGGCCCCGUUAAAUGAGCAGUUAACAUUGCCCGUUAUGAACUUUAAUAUUAAUAACCAUUUAUGCUUAUGACAGAC .((((((((.((((...(.(((((.......))))).)(((.((((.(((((....))))).))))...)))................)))).))))..)))).... (-20.51 = -21.03 + 0.52)

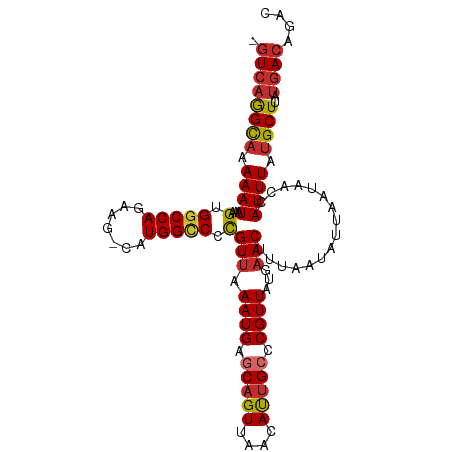
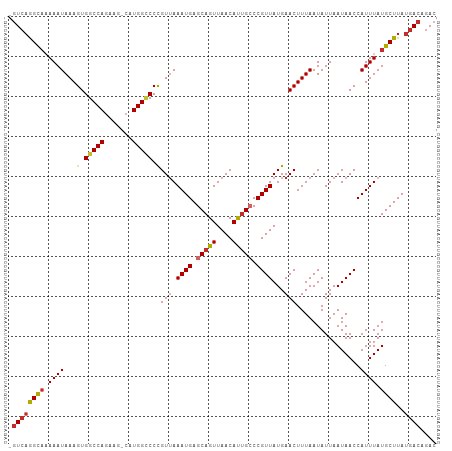
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:37:02 2011