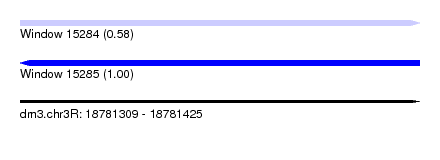

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 18,781,309 – 18,781,425 |
| Length | 116 |
| Max. P | 0.997922 |

| Location | 18,781,309 – 18,781,425 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 94.04 |
| Shannon entropy | 0.11626 |
| G+C content | 0.41090 |
| Mean single sequence MFE | -28.50 |
| Consensus MFE | -23.62 |
| Energy contribution | -25.12 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.83 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.581800 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
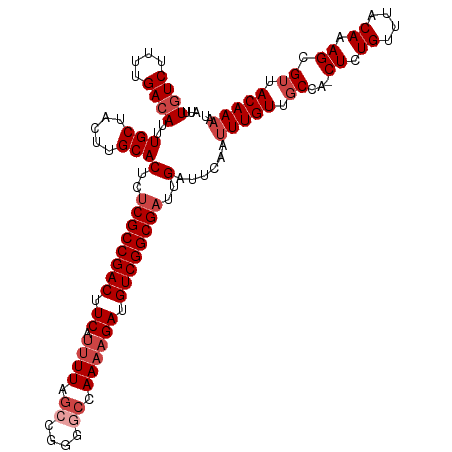
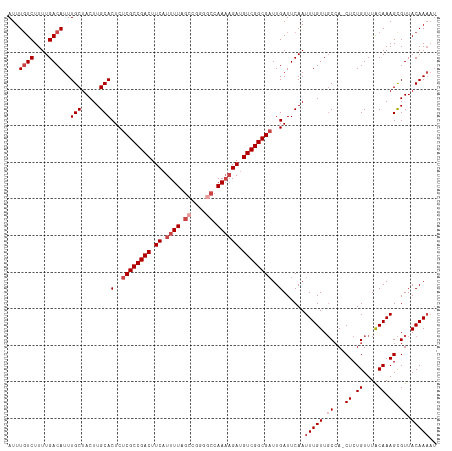
>dm3.chr3R 18781309 116 + 27905053 AUUUGUCUUUUGACAUUUGCUACUUGCACUCUCGCCGACUUCAUUUUAGCCGGGGCCAAAAGAUGUCGGCGAUUGAUUCAAUUUGUUGCCA-CUCUGUUUACAAAGCGUUACAAAAU ...((((....))))..(((.....)))(..((((((((.((.((((.((....)).)))))).))))))))..)......(((((.((..-((.((....)).)).)).))))).. ( -30.70, z-score = -2.29, R) >droSim1.chr3R 18594101 116 + 27517382 AUUUCUCUUUUGACAUUUGCUACUUGCACUCUCGCCGACUUCAUUUUAGCCGGGGCCAAAAGAUGUCGGCGAUUGAUUCAAUUUGUUGCCA-CUCUGUUUACAAAGCGUUACAAAAU .........((((....(((.....)))(..((((((((.((.((((.((....)).)))))).))))))))..)..))))(((((.((..-((.((....)).)).)).))))).. ( -28.60, z-score = -1.98, R) >droSec1.super_0 19145423 116 + 21120651 AUUUGUCUUUUGACAUUUGCUACUUGCACUCUCGCCGACUUCAUUUUAGCCGGGGCCAAAAGAUGUCGGCGAUUGAUUCAAUUUGUUGCCA-CUCUGUUUACAAAGCGUUACAAAAU ...((((....))))..(((.....)))(..((((((((.((.((((.((....)).)))))).))))))))..)......(((((.((..-((.((....)).)).)).))))).. ( -30.70, z-score = -2.29, R) >droYak2.chr3R 19639319 116 + 28832112 AUUUGUCUUUUGACAUUUGCUACUUGCACUCUCGCCGACUUCAUUUUAGCCGGGGCCAAAAGAUGUCGGCGAUUGAUUCAAUUUGUUGCCA-CUCUGUUUACAAAGCGUUACAAAAU ...((((....))))..(((.....)))(..((((((((.((.((((.((....)).)))))).))))))))..)......(((((.((..-((.((....)).)).)).))))).. ( -30.70, z-score = -2.29, R) >droEre2.scaffold_4820 1182165 115 - 10470090 AUUUGUCUUUUGACAUUUGCUACUUGCACUCUCGCCGACUUCAUUUUAGCCGGG-CCAAAAGACGUCGGCGAUUGAUUCAAUUUGUUGCCA-CUCUGUUUACAAAGCGUUACAAAAU ...((((....))))..(((.....)))(..((((((((.((.((((.((...)-).)))))).))))))))..)......(((((.((..-((.((....)).)).)).))))).. ( -27.70, z-score = -1.67, R) >droAna3.scaffold_13340 11157355 112 - 23697760 AUUUGUCUUUUGACAUUUGCUACUUGCACUCUCGCCGACUUCAUUUUUAGCAAGAGCAA--GAUGUCGGCGCUUGAUUCAAUUUGUUCCAAACUCUGUUUGCAAAGUGUUACAA--- ...((((....))))..(((.....))).((.(((((((.....((((.((....))))--)).)))))))...))......((((.....(((.((....)).)))...))))--- ( -22.60, z-score = 0.03, R) >consensus AUUUGUCUUUUGACAUUUGCUACUUGCACUCUCGCCGACUUCAUUUUAGCCGGGGCCAAAAGAUGUCGGCGAUUGAUUCAAUUUGUUGCCA_CUCUGUUUACAAAGCGUUACAAAAU ...((((....))))..(((.....)))(..((((((((.((.((((.((....)).)))))).))))))))..)......(((((.((...((.((....)).)).)).))))).. (-23.62 = -25.12 + 1.50)

| Location | 18,781,309 – 18,781,425 |
|---|---|
| Length | 116 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 94.04 |
| Shannon entropy | 0.11626 |
| G+C content | 0.41090 |
| Mean single sequence MFE | -38.17 |
| Consensus MFE | -30.72 |
| Energy contribution | -32.22 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -4.21 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.21 |
| SVM RNA-class probability | 0.997922 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
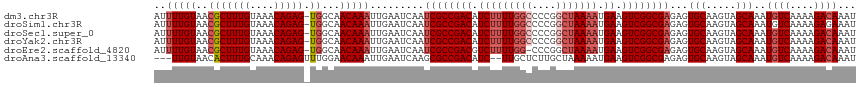
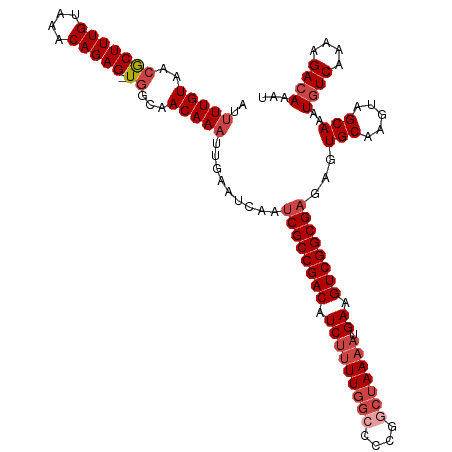
>dm3.chr3R 18781309 116 - 27905053 AUUUUGUAACGCUUUGUAAACAGAG-UGGCAACAAAUUGAAUCAAUCGCCGACAUCUUUUGGCCCCGGCUAAAAUGAAGUCGGCGAGAGUGCAAGUAGCAAAUGUCAAAAGACAAAU ..(((((..(((((((....)))))-))...))))).........((((((((.(((((((((....))))))).)).))))))))...(((.....)))..((((....))))... ( -41.10, z-score = -5.09, R) >droSim1.chr3R 18594101 116 - 27517382 AUUUUGUAACGCUUUGUAAACAGAG-UGGCAACAAAUUGAAUCAAUCGCCGACAUCUUUUGGCCCCGGCUAAAAUGAAGUCGGCGAGAGUGCAAGUAGCAAAUGUCAAAAGAGAAAU ..(((((..(((((((....)))))-))...)))))((((.....((((((((.(((((((((....))))))).)).))))))))...(((.....)))....))))......... ( -39.80, z-score = -4.69, R) >droSec1.super_0 19145423 116 - 21120651 AUUUUGUAACGCUUUGUAAACAGAG-UGGCAACAAAUUGAAUCAAUCGCCGACAUCUUUUGGCCCCGGCUAAAAUGAAGUCGGCGAGAGUGCAAGUAGCAAAUGUCAAAAGACAAAU ..(((((..(((((((....)))))-))...))))).........((((((((.(((((((((....))))))).)).))))))))...(((.....)))..((((....))))... ( -41.10, z-score = -5.09, R) >droYak2.chr3R 19639319 116 - 28832112 AUUUUGUAACGCUUUGUAAACAGAG-UGGCAACAAAUUGAAUCAAUCGCCGACAUCUUUUGGCCCCGGCUAAAAUGAAGUCGGCGAGAGUGCAAGUAGCAAAUGUCAAAAGACAAAU ..(((((..(((((((....)))))-))...))))).........((((((((.(((((((((....))))))).)).))))))))...(((.....)))..((((....))))... ( -41.10, z-score = -5.09, R) >droEre2.scaffold_4820 1182165 115 + 10470090 AUUUUGUAACGCUUUGUAAACAGAG-UGGCAACAAAUUGAAUCAAUCGCCGACGUCUUUUGG-CCCGGCUAAAAUGAAGUCGGCGAGAGUGCAAGUAGCAAAUGUCAAAAGACAAAU ..(((((..(((((((....)))))-))...))))).........((((((((.((((((((-(...))))))).)).))))))))...(((.....)))..((((....))))... ( -38.10, z-score = -3.94, R) >droAna3.scaffold_13340 11157355 112 + 23697760 ---UUGUAACACUUUGCAAACAGAGUUUGGAACAAAUUGAAUCAAGCGCCGACAUC--UUGCUCUUGCUAAAAAUGAAGUCGGCGAGAGUGCAAGUAGCAAAUGUCAAAAGACAAAU ---((((.((((((((....))))))..........(((..((...(((((((.((--..((....)).......)).))))))).))...))))).)))).((((....))))... ( -27.80, z-score = -1.34, R) >consensus AUUUUGUAACGCUUUGUAAACAGAG_UGGCAACAAAUUGAAUCAAUCGCCGACAUCUUUUGGCCCCGGCUAAAAUGAAGUCGGCGAGAGUGCAAGUAGCAAAUGUCAAAAGACAAAU ..(((((.((.(((((....)))))...........(((......((((((((.(((((((((....))))))).)).)))))))).....))))).)))))((((....))))... (-30.72 = -32.22 + 1.50)
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:36:50 2011