
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 18,764,362 – 18,764,455 |
| Length | 93 |
| Max. P | 0.635938 |

| Location | 18,764,362 – 18,764,455 |
|---|---|
| Length | 93 |
| Sequences | 7 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 71.07 |
| Shannon entropy | 0.53642 |
| G+C content | 0.42580 |
| Mean single sequence MFE | -17.51 |
| Consensus MFE | -11.48 |
| Energy contribution | -12.44 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.18 |
| Mean z-score | -0.63 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.635938 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 18764362 93 + 27905053 GUACGACGACUCCAG---CCUCACGUUUGCUGGUAUUUA--AACAAUCUAAAUGAUGCUCUUGGUACUGCCAGCAAAUUGAAAUUUACUGCCCCUCUU------------------ ...............---..(((.(((((((((((.(((--(...(((.....)))....))))...)))))))))))))).................------------------ ( -17.30, z-score = -1.15, R) >droSim1.chr3R 18575625 93 + 27517382 GUACGACGACUCCAG---CCUCACGUUUGCUGGUAUUUA--AACAAUCUAAAUGAUGCUCUUGGUCCUGCCAGCAAAUUGAAAUUUACUGCCCCUCUU------------------ ...............---..(((.(((((((((((.(((--(...(((.....)))....))))...)))))))))))))).................------------------ ( -17.30, z-score = -1.17, R) >droSec1.super_0 19128838 93 + 21120651 GUACGACGACUUCAG---CCUCACGUUUGCUGGUAUUUA--AACAAUCUAAAUGAUGCUCUUGGUCCUGCCAGCAAAUUGAAAUUUACUGCCCCUCUU------------------ ...............---..(((.(((((((((((.(((--(...(((.....)))....))))...)))))))))))))).................------------------ ( -17.30, z-score = -1.10, R) >droYak2.chr3R 19622034 93 + 28832112 GUACGACGACUCCAG---CCUCACGUUUGCUGGUAUUUA--AACAAUCUAAAUGAUGCUCUUGGUCCUGCCAGCAAAUUGAAAUUUACUGCCCCUCUU------------------ ...............---..(((.(((((((((((.(((--(...(((.....)))....))))...)))))))))))))).................------------------ ( -17.30, z-score = -1.17, R) >droEre2.scaffold_4820 1165867 90 - 10470090 GUAGGACGACUCCAG---CCUCACGUUUGCUGGUAUUUA--AACAAUCUAAAUGAUGCUCU-GGUCCUGCCAGCAAAUUGAAAUUUACUGCCCCUU-------------------- ((((((....)))..---..(((.(((((((((((....--....(((.....))).....-.....))))))))))))))....)))........-------------------- ( -19.67, z-score = -1.21, R) >droAna3.scaffold_13340 11139999 107 - 23697760 ---CUCGGACUCCU----UCUCAGUUUUGCUGGUAUUUA--AACAAUCUAAAUGAUGCUUUCGGUUCUGGCUGCAAAUUGAAAUUUACUGCCGUCUCCCCGCCUCUUCCGGUCUGA ---.(((((((...----..(((((((.((..(.((..(--((..(((.....)))..)))..)).)..))...)))))))........((.(.....).)).......))))))) ( -19.20, z-score = 0.04, R) >droWil1.scaffold_181089 11102094 113 - 12369635 GUUUGGCAAAUUUUGGCAUUGCAUAUCUGUUUUUCUUUGUUGUCAUUCCUCUUGGUAUGUCUAGUUUUG-CUACAAAUUGAAAUUUACUACUGUCGACUGGAUAACAUUUUUGG-- (..((((((.....((.(..(((....)))..).))...))))))..)......((.(((((((((..(-(.....................)).)))))))))))........-- ( -14.50, z-score = 1.32, R) >consensus GUACGACGACUCCAG___CCUCACGUUUGCUGGUAUUUA__AACAAUCUAAAUGAUGCUCUUGGUCCUGCCAGCAAAUUGAAAUUUACUGCCCCUCUU__________________ ....................(((.(((((((((((..........(((.....)))...........))))))))))))))................................... (-11.48 = -12.44 + 0.96)
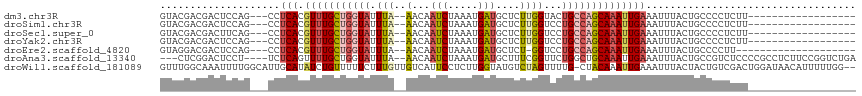

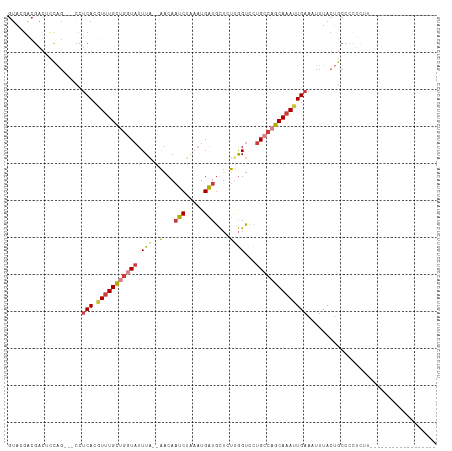
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:36:41 2011