
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 18,752,591 – 18,752,691 |
| Length | 100 |
| Max. P | 0.837328 |

| Location | 18,752,591 – 18,752,691 |
|---|---|
| Length | 100 |
| Sequences | 8 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.53 |
| Shannon entropy | 0.43459 |
| G+C content | 0.38719 |
| Mean single sequence MFE | -27.77 |
| Consensus MFE | -14.75 |
| Energy contribution | -14.76 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.54 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.86 |
| SVM RNA-class probability | 0.837328 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
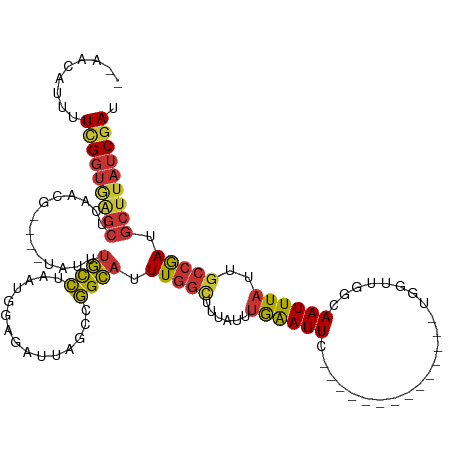
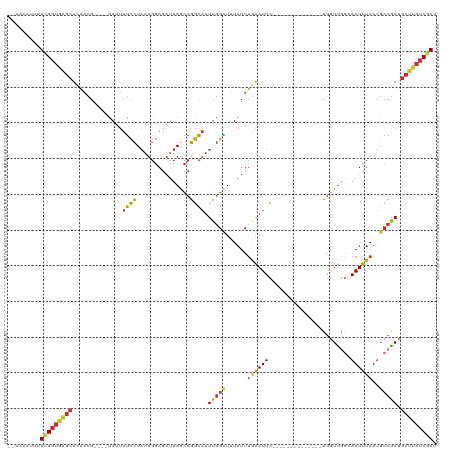
>dm3.chr3R 18752591 100 + 27905053 --AACAUUUUCGGUGAGCUCAACG----UAUUUGCCUAAUGGAGAUUAGCCGGCAUUUGGCUUUAUUUGAAUUC--------------UGGUUGGCAAUUUAUUGCCAAUGCUUAUCGAU --.......(((((((((((((.(----((...(((.((((..(......)..)))).)))..)))))))....--------------..((((((((....))))))))))))))))). ( -31.30, z-score = -2.98, R) >droSim1.chr3R 18563667 100 + 27517382 --AACAUUUUCGGUGAGCUCAACG----UAUUUGCCUAAUGGAGAUUAGCCGGCAUUUGGCUUUAUUUGAAUUC--------------UGGUUGGCAAUUUAUUGCCGAUGCUUAUCGAU --.......(((((((((((((.(----((...(((.((((..(......)..)))).)))..)))))))....--------------..((((((((....))))))))))))))))). ( -31.00, z-score = -2.76, R) >droSec1.super_0 19117377 100 + 21120651 --AACAUUUUCGGUGAGCUCAACG----UAUUUGCCUAAUGGAGAUUAGCCGGCAUUUGGCUUUAUUUGAAUUC--------------UGGUUGGCAAUUUAUUGCCGAUGCUUAUCGAU --.......(((((((((((((.(----((...(((.((((..(......)..)))).)))..)))))))....--------------..((((((((....))))))))))))))))). ( -31.00, z-score = -2.76, R) >droYak2.chr3R 19609862 100 + 28832112 --AACAUUUUCGGUGAGCUCAACG----UAUUUGCCUAAUGGAGAUUAGCCGGCAUUUGGCUUUAUUUGAAUUC--------------UGGUUGGCAAUUUAUUGCCGAUGCUUAUCGAU --.......(((((((((((((.(----((...(((.((((..(......)..)))).)))..)))))))....--------------..((((((((....))))))))))))))))). ( -31.00, z-score = -2.76, R) >droEre2.scaffold_4820 1154224 100 - 10470090 --AGCAUUUUCGGUGGGCUCAACG----UAUUUGCCUAAUGGAGAUUAGCCGGCAUUUGGCUUUAUUUGAAUUC--------------UGGUUGGCAAUUUAUUGCCGAUGCUUAUCGAU --.......(((((((((((((.(----((...(((.((((..(......)..)))).)))..)))))))....--------------..((((((((....))))))))))))))))). ( -30.20, z-score = -1.75, R) >droAna3.scaffold_13340 11128196 112 - 23697760 ----AAAAUUCGGUAAGCUCAUCG----UAUUUGCCUAAUGGAGAUUAGCCGGCAUUUGGUUUUAUUUGAAUUCAGGGCCGUGCACGGUGGUCGGAAAUUUAUUGCCGAUGCUUAUCGAU ----.....(((((((((.(((((----(((((.((....))))))..((((((.((((..(((....)))..)))))))).))))))))(((((..........)))))))))))))). ( -37.60, z-score = -2.71, R) >droWil1.scaffold_181089 11089597 113 - 12369635 AAUACAUUUUUGGUAAGCUGAACGAACGCAUUUGCCUAAUGGAGAUUAGCCGGCAUUUGGUUUUAUUUGAAUU-----UCAUCCGUGGU--UUCCAAAUUUAUUGCCGAUGCUUAUCGAU .........(((((((((.........((((((.((....))))))..))((((((((((..((((.((....-----.))...)))).--..))))).....)))))..))))))))). ( -24.00, z-score = -0.17, R) >droGri2.scaffold_15074 3556779 94 + 7742996 --AGCUUUCUCGCAUAUCUUGAUG----AACUUUUUUUCCCCACUUUAAACAGACUUUUGAUUUAUUUAGAUUU--------------UGAUU---AAUUUUU---CAACACAUGUCAAC --.((......)).....((((((----((......)))......((((.((((..(((((.....))))).))--------------)).))---)).....---........))))). ( -6.10, z-score = 0.75, R) >consensus __AACAUUUUCGGUGAGCUCAACG____UAUUUGCCUAAUGGAGAUUAGCCGGCAUUUGGCUUUAUUUGAAUUC______________UGGUUGGCAAUUUAUUGCCGAUGCUUAUCGAU .........(((((((((((((......((..((((...............))))..)).......))))....................(((((..........)))))))))))))). (-14.75 = -14.76 + 0.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:36:36 2011