| Sequence ID | dm3.chr3R |
|---|---|
| Location | 18,712,300 – 18,712,392 |
| Length | 92 |
| Max. P | 0.940796 |

| Location | 18,712,300 – 18,712,392 |
|---|---|
| Length | 92 |
| Sequences | 10 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 68.22 |
| Shannon entropy | 0.64972 |
| G+C content | 0.39212 |
| Mean single sequence MFE | -20.51 |
| Consensus MFE | -10.76 |
| Energy contribution | -10.77 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.19 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.47 |
| SVM RNA-class probability | 0.940796 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 18712300 92 - 27905053 -----AGCUGCAUUGAUGGUCUGGGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCAGUUUCAGUUUUCAUUUG--CCCCCUUUUUAUUUGUUGU--- -----.((.(((.(((.((...(((((((((((((((((((.(((((.....)))))))))))))))).....)))))--)))))...)))..))).))--- ( -28.20, z-score = -2.93, R) >droSec1.super_0 19077592 90 - 21120651 -----AGCUGCAUUGGUGGGCUGAGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCAGUUUCAGUUUUCAUUUG---CGCCUUUUUA-UUGUUGU--- -----.((.(((.(((.((((...(((((((((((((((((.(((((.....)))))))))))))))).....)))))---)))))..)))-.))).))--- ( -27.10, z-score = -1.97, R) >droYak2.chr3R 19566043 77 - 28832112 ----------------------AAGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCCGUUUCAGUUUUCAUUUGGCCCCCUUUUUUAUUUGUUGU--- ----------------------.((((((((((((((.(((.(((((.....)))))))).))))))).........((...))......)))))))..--- ( -13.80, z-score = -0.62, R) >droEre2.scaffold_4820 1114180 91 + 10470090 -----GGCUGCAUUGGUGGGCUGAGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCAGUUUCAGUUUUCAUUUG---CCCCCUUUUAUUUGUUGU--- -----.((.(((..((.((((.((......(((((((((((.(((((.....))))))))))))))))......)).)---))))).......))).))--- ( -27.10, z-score = -1.85, R) >droAna3.scaffold_13340 11087261 80 + 23697760 ----------------------UGGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCAGUUUCAGUUUUCAUUCCGCUUGUCCGUUUUCCUUUUUUUCU ----------------------.(((((...((((((((((.(((((.....)))))))))))))))(....).......)))))................. ( -16.00, z-score = -1.40, R) >dp4.Unknown_group_570 804 82 - 14025 ------------GUUGUGGGCUUGGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCCGUUUCAGUAUCCUGUU---UCUUUUUUCUUUCUGUC----- ------------.....((((..((.....(((((((.(((.(((((.....)))))))).)))))))...)).)))---)................----- ( -13.00, z-score = 0.45, R) >droPer1.super_145 74734 82 + 90410 ------------GUUGUGGGCUUGGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCCGUUUCAGUAUCCUGUU---UCUUUUUUCUUUCUGUC----- ------------.....((((..((.....(((((((.(((.(((((.....)))))))).)))))))...)).)))---)................----- ( -13.00, z-score = 0.45, R) >droWil1.scaffold_181089 11046973 93 + 12369635 AGGACAAAGGUCAUCUUGGGCUUGGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCAGCUUUAGGUUCAGUUUAUUUUUCCAUCUUUGU--------- ...((((((((.....((((((.((...(((((((.(((((.(((((.....)))))))))).))))))))))))))).......))))))))--------- ( -19.20, z-score = -0.23, R) >droVir3.scaffold_12822 3924810 99 + 4096053 ---CUUUUUGUUGCUUUUUGCUCGGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCAGUUUCAGUUUCCAUUUCGUUUGUCCAUUUGCCAUUUGGUCU ---.........((.....))..((((((((((((((((((.(((((.....))))))))))))))))....((.......))...)))))))......... ( -23.20, z-score = -1.78, R) >droMoj3.scaffold_6540 1244342 93 + 34148556 ---------CUUGGUUUUGCUUUGGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCAGUUUCAGUUUCCAUUUCGUUUGCCCAUUCGCCAUUUGGUUU ---------..((((..((....((((((((((((((((((.(((((.....))))))))))))))))..........)))))))))...))))........ ( -24.50, z-score = -1.96, R) >consensus _________G__GUUGUGGGCUUGGCAAAUUUGAAAUUGAUAUUCGCACGGCGUGAAAUCAGUUUCAGUUUCCAUUUG__UCGCCUUUUUAUCUGUUGU___ ..............................(((((((.(((.(((((.....)))))))).))))))).................................. (-10.76 = -10.77 + 0.01)
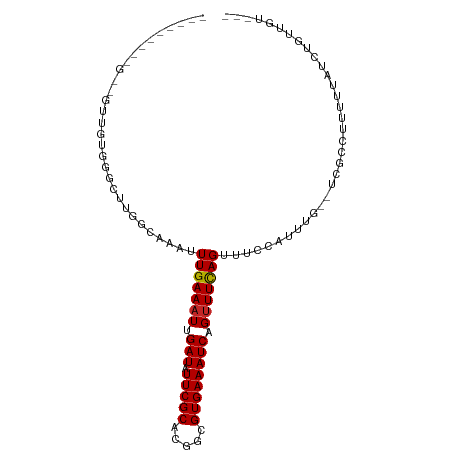


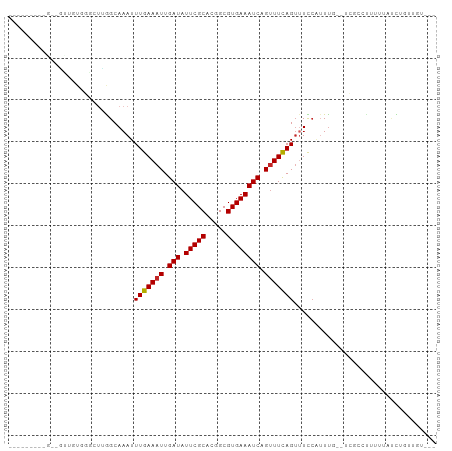
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:36:23 2011